Possible Therapeutic Targets from Derivatives of Natural Marine Products Based on PI3K/AKT Dependent Inhibitors in Viral Infection COVID-19
bNutritional Health Team (NHT), Universal Scientific Education and Research Network (USERN), Tehran, Iran,
cMedical Plants Research Center, Basic Health Sciences Institute, Shahrekord University of Medical Sciences, Shahrekord, Iran,
dDepartment of Cancer Biology, Lerner Research Institute, Cleveland Clinic, Cleveland, OH, USA,
eUniversity of Florida Health Cancer Center, Gainesville, FL, USA
fUniversity of Florida, College of Nursing, Gainesville, FL, USA,
gFaculty of Medicine, Shahid Sadoughi University of Medical Sciences, Yazd, Iran,
hFaculty of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran,
iShahrekord University of Medical Sciences, Shahrekord, Iran,
jVascular and Endovascular Surgery Research Center, Mashhad University of Medical Science, Mashhad, Iran
Keywords
Abstract
Natural resources have long played a prominent part in conventional treatments as a parental source due to their multifaceted functions and lesser side effects. The diversity of marine products is a significant source of possible bioactive chemical compounds with a wide range of potential medicinal applications. Marine organisms produce natural compounds and new drugs with unique properties are produced from these compounds. A lot of bioactive compounds with medicinal properties are extracted from marine invertebrates, including Peptides, Alkaloids, Terpenoids, Steroids. Thus, it can be concluded that marine ecosystems are endowed with natural resources that have a wide range of medicinal properties, and it is important to examine the therapeutic and pharmacological capabilities of these molecules. So, finding particular inhibitors of the COVID-19 in natural compounds will be extremely important. Natural ingredients, in this light, could be a valuable resource in the progression of COVID-19 therapeutic options. Controlling the immunological response in COVID-19 patients may be possible by addressing the PI3K/Akt pathway and regulating T cell responses. T cell effector activity can be improved by preventing anti-viral exhaustion by suppressing PI3K and Akt during the early anti-viral response. The diversity of marine life is a significant supply of potentially bioactive chemical compounds with a broad range of medicinal uses. In this study, some biologically active compounds from marine organisms capable of inhibiting PI3K/AKT and the possible therapeutic targets from these compounds in viral infection COVID-19 have been addressed.
Introduction
Due to the accelerating spread of the COVID-19 pandemic and the considerable time required to develop vaccines and effective drugs, a pressing need exists to develop strategies for preventing and treating COVID-19 using the already existing natural products [1-3]. Numerous natural products have been widely studied and used as potential options for the treatment of COVID-19 [4-6]. It seems that the endosomal–autophagic vesicles networks play a key role in the infections caused by Coronaviruses (CoVs), including SARS-CoVs [7, 8]. Autophagy plays an important role in pulmonary infections, improving the host’s immune defenses against bacterial and viral infections of the respiratory tract. Considering the various roles of autophagy in viral infection, it was postulated that three groups of autophagy modulators might inhibit viral replication clinically related to COVID-19 [9, 10]. The first group is the drugs with lysosomotropic activity, which inhibit cathepsin activity, preventing infections caused by CoVs by neutralizing the acidic pH in the endosomes-lysosomes [8]. The second group of drugs includes protease inhibitors, which might inhibit the S protein’s proteolytic cleavage and limit viral cell entry. The third group of drugs includes PI3K/AKT/mTOR regulators, which might mean autophagic machinery’s CoVs-mediated appropriation, despite being considered autophagy regulators [11]. In the next subsections, several well-tolerated clinically approved autophagy-modulating compounds will be discussed, which might be investigated as potential COVID-19 modulators and replication used in COVID-19 management. Currently, there is not any acceptable antiviral drug to treat SARS-CoV-2 infections. Hence, preventive and supportive cares of complications are essential strategies to minimize the harms [11, 12]. Suggested strategies for the treatment of SARS-CoV-2 might be divided into four different categories: SARS-CoV-2 RNA synthesis prevention, through vital proteins and enzymes inhibition, virus-cell receptor binding impedition or virus self-assembly suppletion, the host’s immunity stimulation by producing virulence factors, and virus entry blockage into host cells by acting on the host’s enzymes or through cellular entry receptors [13-16]. Alkaloids, macrolides, polypeptides, and terpenoids are just a few of the molecular classes represented by marine-derived natural products and contain a variety of unique and fascinating structures that contribute significantly to biological activity and clinical therapeutic uses. In terms of kinase research, several marine-derived kinase inhibitors have been developed from a variety of various sources and have been shown to inhibit a wide range of protein kinases. Recently, several biologically active kinase inhibitors have been introduced in different ocean life forms, such as soft corals, algae, fungi, bacteria, cyanobacteria, sponges, and animals. Basic bioactivity tests are usually included in the initial reports of these natural agents, and subsequent researches should go on to evaluated the pharmacological activity of each in greater depth [17-20].
Lipid Kinase Inhibitors
Phosphoinositide 3-Kinase (PI3K) Inhibitors
PI3K, belonging to a large family of lipid kinases, has been proven to play a significant role in cancer, aging, and diabetes. PI3Ks have a “PI3K characteristic motif”, composed of a C2 domain (which may bind to the membrane), a catalytic kinase domain, and a helical domain [21, 22]. There are three types of PI3K (I, II, III) and four isoforms (α, β, γ, δ). Class I (I PI3K) has been implicated in various cancers [22, 23].
Class I PI3Ks.
Class I PI3K is a heterogeneous mer, including coordination (p50α, p55α, p55γ, p85α, p85β, or p101) subunits with catalysts (p110α, p110β, p110δ, or p110γ). The known carcinogenic PI3K/mTOR signaling has been explored elsewhere [24]. In outline, this pathway is actuated by cell surface receptors, such as oncogenes like RAS, G protein-coupled receptors, and receptor tyrosine kinases. Furthermore, the activated p110 subunit can catalyze converting phosphatidylinositol 4,5-bisphosphate (PIP2) to phosphatidylinositol 3,4,5-trisphosphate (PIP3), activating protein kinase B/AKT. AKT usually sends signals to downstream effectors, such as glycogen synthase kinase 3, forkhead transcription factor, mTOR complex (mTORC) 1, and cell death antagonist BCL2 to regulate different cellular processes. mTORC2 also promotes AKT activation through the serine 473 phosphorylation. Contrasting to this, the role of phosphotensin homolog (PTEN) is antagonizing the PI3K function and dephosphorylating PIP3 to PIP2 [24, 25]. Many growth factor pathways are controlled by activated RTK or GPCR. These RTKs or GPCRs recruit the p85p110 complex to the plasma membrane to alleviate the inhibition of p85, and p110 converts PtdIns (4,5) P2 to PtdIns (3,4,5) P3 to trigger a signal conduction response [25, 26]. In particular, PtdIns (3,4,5) P3 recruits Akt kinase, which controls the activation of FOXO, mTORC1, etc. to regulate metabolism, autophagy, survival, cell growth, and proliferation [27, 28]. Restricted class I PI3K movement additionally assumes a part in cortical F-actin elements, which causes chemotaxis and phagocytosis of particles [29-32]. For instance, at the neutrophil driving edge, p110γ-incited PtdIns (3,4,5) P3 development brings about the enrollment of Rac GTPase, which advances F-actin polymerization, lamellipodia arrangement, and cell relocation [33].
3phosphatase PTEN downregulates PtdIns (3,4,5) [3-5] P3 and class I PI3K activation pathways [34]. Type 4 I catalyst isomorphs are nested but share distinct functions. While p110γ and p110δ isomorphs are primarily restricted to functions expressed in immune cells, p110α and p110β also exhibit isomorphic-specific cell types and context-dependent requirements. Most Class I PI3K functions are associated with catalytic properties. However, there is increasing evidence of the role of kinase-independent scaffolding of p110γ and p110β [35-37].
Class II PI3Ks.
Class II PI3K includes Class I and Class III and other high molecular weight monomers with long N- and C-terminal domains [38]. The magnificence II enzymes lack structural records that could assist in recognizing those mechanisms. While the magnificence II enzymes include a PI3K center and a Ras-binding area (RBD) like different individuals of the PI3K family, all magnificence II enzymes include C-terminal domain names, along with a phox-homology (PX) area observed with the aid of using a C2 area. A prolonged N-terminal area is particular to magnificence II. Unlike their magnificence I and III spouse and children, which can be focused on their web website online of movement through special regulatory subunits, magnificence II enzymes lack solid affiliation with regulatory subunits. Instead, those particular domain names at each terminus are idea to play an essential position inside the law and localization of those kinases with the aid of using mediating particular protein and lipid interactions [39]. In mammals, 3 totally different category II members are identified: the ubiquitously expressed PI3K-C2 α, the liver-specific PI3K-C2 γ, and PI3KC2 β [38, 40]. Contrasting to class I, class II PI3Ks are primarily recognized as substrates PtdIns and PtdIns (4)P, which are involved in the regulation of follicular transport, resulting in PtdIns (3) P and PtdIns (3,4) P2, respectively. Several stimuli, including cytokines, chemokines, growth factors, and hormones can activate class II PI3Ks through various membrane receptors, such as GPCRs and RTKs (EGFR and PDGFR) [38-41]. The best studied in class IIPI3K is PI3KC2α. At the cellular level, it is demonstrated that PI3KC2α regulates the complete translocation of the glucose transporter GLUT4 to the plasma membrane of muscle cells, an important event in the regulation of glucose homeostasis [41].
Class III PI3Ks.
PI3K is a lipid kinase that can phosphorylate phosphatidylinositol (PI) and its phosphorylated derivatives phosphatidylinositol 4,5 diphosphate (PIP2) and phosphatidylinositol 4 The D3 hydroxyl group of monophositol (PI4 ring). PI3K accepts entry from activated receptor tyrosine kinases and heterotrimeric guanine nucleotide (G protein) -binding protein-coupled receptors (GPCRs), and affects cell proliferation, growth, metabolism, motility, and second messenger transport of intracellular lipids [42]. In recent years, mammalian class III PI3K (phosphatidylinositol 3 kinase) complex has become the key to several basic cellular processes through the aftereffect of its catalytic product phosphatidylinositol 3-phosphate (PtdIns (3) P) Modifier. One of these processes is the ligand-dependent down-regulation of growth factor receptors through membrane transport events. Specifically, PtdIns (3) P mediates the load classification from early endosomes to multivesicular bodies and finally degraded lysosomes [43]. Inhibition of mTOR results in the activation of the important molecule Unc51like autophagy effective kinase 1 (ULK1), which is potentiated into the endoplasmic reticulum (ER) for the initiation of autophagy. ULK1 is a crescent-shaped bilayer structure known for omega cotton that emerged from the ER by recruiting additional III-type four-spoy city Ted 3 kinase (PI3K) complexes composed of Beclin1, VPS34, and ATG14. Induces formation and macrophage nucleation. The class III PI3K complex mediates the accumulation of phosphatidylinositol triphosphate (PIP3) on the surface of omega cotton to recruit LC3, and phosphatidylethanolamine (PE) is conjugated to LC3I to form LC3II, which forms a phagocytic membrane. Phagocytes eventually seal, form self-predatory bodies, and then fuse with lysosomes to break down the packaged contents. Apart from recycling useful substances, self-predation is also an important mechanism for removing intracellular pathogens, including viruses, which is called virophage [43-45].
Immune system and activation of PI3K
Unique PI3K lineage members are active in the immune system, depending on the type of receptor and/or cell [41, 46-48]. For instance, cytokines like interleukin (IL)-2, IL-3, IL-6, IL-7, and IL-15, interferons (IFNs), oncostatin M, erythropoietin, granulocyte colony-stimulating factor activate class IA PI3Ks in a number of immune cells, such as dendritic cells (DCs) and T cells, by activating Janus kinase and tyrosine phosphorylation of different proteins. Cytokine receptors with intrinsic PTK activity can also activate class IA PI3Ks. These enzymes are further activated on antigen recognition through natural killer (NK) stimulatory receptors, the B cell receptor (BCR), T cell receptor (TCR), and Fc receptors, the same a IgG (FcγRI) and high-affinity IgE (FcεRI) receptors [49-52]. Co-stimulatory receptors such as CD28 and CD19 on T cells and B cells, respectively, and cell adhesion molecules also activate class IAPI3Ks. The cytoplasmic region of CD28 and CD19 binds to the p85 subunit, so binding of both antigen receptors and costimulatory receptors potently activates PI3K [52-54]. Various adapter proteins including BCAP and linkers for T cell (LAT) activation that mediate protein-protein interactions are involved in the activation of IAPI3K, a downstream class of antigen and costimulatory receptors, possibly recruited to the plasma membrane of tyrosine phosphorylation [53-55]. In particular, the tumor necrosis factor receptor family, such as the IL1 receptor Toll-like receptor (TLR) and CD40, activate class IAPI3Ks in many cell types, including macrophages and DCs, despite no apparent association with PTK [41, 46, 53, 54]. It is not yet clear how signal transduction through these receptors enables downstream PI3K. Unlike class IA PI3Ks, class IB PI3Ks are mostly activated via GPCRs like chemokine receptors [56-59]. Class II and class III PI3Ks are likely to play a role in the immune system, but the mechanisms supporting their activation remain undetermined. Giant predation, the process by which long-lived proteins and intracellular components such as organelles are broken down and recycled, is at least partially regulated by the PI3KAKT path [58-60]. I It can be additionally induced by activating kinase Ulk1 throu AMP-activated protein kinase (AMPK). Activating the PI3KAKT mTORC1 path suppresses self-predation through suppression of the interaction between AMPK and Ulk1 by mTORC1 [61]. Auto-purging with respect to viral infections has been well-studied, but it is not necessarily clear whether the effectiveness of the route is pro-virus or anti-virus [62, 63]. The advantages of autophagy appear to be distinct for distinct viruses, depending on the time after infection. Some viruses, inclusive of rotavirus and ZIKV, require the manner early in infection [27, 61-63], while that is unfavorable for others. Conversely, a few viruses, including influenza A virus result in autophagy overdue after contamination to boom replication. Finally, there are viruses wherein autophagy does now no longer seem to steer replication. Autophagy has been studied withinside the context of CHIKV, SFV, and SINV infection, aleven though maximum of those research did now no longer check out the direct position of PI3K-AKT activation. These alphaviruses seem to have special consequences on autophagy, aleven though there additionally is probably version because of special mobileular sorts used and special experimental conditions [63-65].
Akt
Akt, a serine/threonine kinase, which was previously known as protein kinase B (PKB), is composed of Akt1, Akt2, and Akt3 that regulate glucose metabolism, cell cycle progression and cell size. Akt plays an important role in key cellular functions, including neovascularization, protein synthesis, genome, stability, and transcription. Akt facilitates cell survival by blocking apoptosis through the ineffectiveness of pro-apoptosis proteins, which mediates cell growth factors [66-70]. Akt/PKB is similar to supermolecule kinase A (PKA) and kinases C (PKC), further on the retroviral oncoprotein microorganism akt (v akt) [71-74]. Akt consists of a central, an amino terminus (N terminus), and a carboxyl terminus fragment (C terminus). The purecustorin homology (PH) domain (N-terminal domain), consisting of 100 amino acids, is similar to other domains seen in 3-hoshoinoshichido binding molecules that interact with membrane lipid products such as phosphatidylinositol triphosphate (PIP3) and phosphatidylinositol 4,5-diphosphate (PIP2) [68-70]. The kinase domain, being similar to the AGC protein kinase, shares a regulatory residue named Thr308, which its phosphorylation activates Akt. The C-terminal home is composed of forty amino acids that form a hydrophobic region containing the regulatory serine residue Ser473 [71, 72].
PI3K/AKT (Structure and function)
Akt kinase hobby is precipitated following PI3K activation in numerous boom thing receptor-mediated signaling cascades [75]. PI3K phosphorylates hoshoinoschido at the 30OH position of the inositol ring. The secondary messenger products of the kinase response in growth factor-stimulated animal cells are phosphatidylinositol 3,4 phosphate (PI3, 4P2) and phosphatidylinositol 3,4,5 phosphate (PIP3) [76]. The discovery of phosphoinositide-specific phospholipid phosphatases, such as PTEN, further enhances our knowledge due to the regulation of secondary messenger signals by phospholipids and cellular phosphoinositide metabolism. Activation of Akt by PI3K is initiated through binding specific 30 phosphorylated Force Spoi City Ted in the Akt PH domain [77]. The Akt PH area binds to each PIP3, and PI-3,4-P2 and suggests a fairly better affinity for the binding of PI-3,4-P2. The binding specificities of various PH domain names for one of a kind phosphoinositides shown with the aid of using structural studies, and the specificity of the Akt PH area for each PI-3,4-P2 and PIP3 can be applicable for accomplishing a prolonged sign of Akt pastime in cells [76]. The initial binding result of plasma-bound Force Spoiled Ted Akt is the reorganization of the cellular Akt protein of the plasma transduction complex. The PH domain is important for mediating these reorganizations, consistent with its ability to bind to phospholipid second messenger molecules. The discovery of increased activity of the N-terminal deleted Akt mutation was interpreted as an indication of intramolecular inhibition, but only recent evidence suggested an interaction of the N- and C-terminal regions of the Akt molecule [78]. Oncogenic mutations in Akt, which would result in constitutive factor-independent activation, also account for activity in the vakt configuration by involving the plasma membrane binding to the N-terminal myristoyl, thereby circumventing the initial translocation step [79, 80]. Phosphorylation with the aid of using upstream kinases is needed for complete activation of Akt and is essential for protection of its activity. Therefore, it isn’t always sudden that a few protein phosphatases are capable of inactivate Akt with the aid of using dephosphorylation. Nonspecific inhibition of endogenous phosphatase hobby turns on Akt efficiently [81]. Furthermore, phosphatase activity is concerned in mediating the results of extracellular stresses on Akt sign transduction [80]. Akt dephosphorylation was also observed after increasing ceramide levels, possibly due to inhibition of PI3K or other upstream signaling molecules [82]. Information on the importance of phosphatases for the regulation of Akt activity was also obtained from the study of the phosphatases, which dephosphorylated the phosphoinositide products of PI3K. Among these, the homology of the src2 (SH2) domain containing the inositol SHIP1/2 phosphatases and the PTEN phosphatase is important mediators in the determination of Akt activity. SHIP1 dephosphorylates inositides and phosphoinositides at position 50 and regulates B cell / myeloid function [82-84].
PI3K signaling activation
In normal situation, the catalytic subunit (p110) is synthetized through dimerization with the regulatory subunit (p85) and different extracellular stimuli, including cytokines, hormones, and growth factors activate PI3K [85]. When activated, PI3K phosphorylates PtdIns (4,5) P2(PIP2) to make PtdIns (3,4,5) P3(PIP3) and recruits a subset of pleckstrin-homology (PH), such as Phox (PX), FYVE, C1, C2 to the cell membrane. Several signaling proteins, including kinases PDK1 and AKT, may localize to the cell membrane by binding to the PI3K lipid products to activate cell growth and survival [86]. The pathway is normally regulated through dephosphorylating PIP3 to PIP2 and preventing the downstream kinases’ activation [87].
PI3K and SARS-CoV-2
PI3K/AKT signaling plays a crucial role in migration, invasion, cell proliferation, growth, and survival, and can promote angiogenesis and subset cell apoptosis. Abnormal PI3K/AKT signaling pathway has been shown to cause health problems and diseases such as cancer. With life science advancements, targeted therapy such as PI3K/AKT inhibition has become a popular method for the treatment of malignant tumors. AKT and PI3K are antitumor drug targets, and their antitumor therapies have shown attractive prospects. A potent panAKT kinase inhibitor, Capivasertib, which inhibits AKT1, AKT2, and AKT3 has shown an acceptable antitumor activity. As an AKT inhibitor, Capivasertib is used in clinical trials for the treatment of drug-resistant breast cancers [88, 89]. The kinase inhibitor capivasertib is also an anticancer drug that targets AKT preventing SARSCoV2 from entering the cell. There are a lot of antitumor drugs that target the PI3K/AKT signaling pathway. This might be used in treating critical COVID19 cancer patients in the era of the pandemic [89-91].
(PI3K)/AKT and development of immune responses
The development of immune responses involves entering the virus into the cell and immune response development. The relationship between virus entery and this pathway should be examined. The main receptors involved in virus entery are and CD147 and ACE2 [91-93]. Furin and TMPRSS2 are two other non-endosomal pathways for viral entry [93]. Furin and CD147activation can induce the PI3K/AKT signaling pathway [93, 94]. Endocytosis occurs when the virus binds to ACE2. The occurrence of SARS-CoV-2 endocytosis is regulated by the PI3K/AKT signaling pathway and is mediated through a clathrin pathway [93-95]. Suppression of this pathway can inhibit the entry of any clathrin-mediated endocytosis. ACE2 reduction at the cell surface enhances the serum levels of Ang II, which is seen in COVID-19 patients [96-99]. Ang II elevation in serum can enhance the inflammatory cytokines such as TNF-α and IL-6, as seen in COVID-19 patients. Ang II also induces the fibrosis of various organs, which can be attributed to the PI3K/AKT signaling pathway [97, 99, 100]. The binding of Ang II to ang II receptor type 1 (AT1R) can activate this pathway [101]. Activation of PI3K/AKT signaling pathway induces lung tissue fibrosis, which is frequently seen in COVID-19 patients [91, 97, 102]. The activation of some factors including nuclear factor kappa B (NF-κB) and activated protein-1 (AP-1) has a crucial role in induction of inflammation [103]. If the PI3K/AKT signaling pathway is suppressed, AP-1 and NF-κB is inhibited reducing the inflammatory cytokines such as TNF-α and IL-6 [91, 104]. Hence, PI3K/AKT inhibitors can suppress inflammation, and their uses along with antiviral drugs help combat COVID-19 [105, 106].
PI3K/AKT signaling in human viral infections
The PI3K/AKT signaling pathway is known to be modulated by many viruses. Most viruses activate the pathway through different measures; therefore, they prevent cell apoptosis, an effective and primitive host defense mechanism [107, 108]. As an essential step to maintain reproduction and dissemination, interruption in programmed cell death enables viruses to enhance replication and even cause persistent infections. Adenoviruses activate PI3K/AKT pathway in corneal and lung epithelial cells, respectively.
Hepatitis B virus (HBV) is known to be an activator of PI3K/AKT cascade [107, 109-112]. On the other hand, HBV DNA replication is downregulated by the active PI3K/AKT signaling pathway, making the latter an equalizer of acute infection and hepatocarcinogenesis [113, 114]. Interestingly some viruses affect the course of other viral infections by the means of PI3K/AKT pathway. Hepatitis C virus (HCV) NS5A protein reduces HBV DNA replication by activating PI3K/AKT cascade [113]. Another example is human immunodeficiency virus (HIV) Tat protein, an activator of PI3K/AKT pathway, which prevents apoptosis. In HIV infected patients Kaposi’s sarcoma-associated herpes virus (KSHV) causes a more aggressive form of Kaposi’s sarcoma, which is believed to be associated with the anti-apoptotic features of Tat protein. It is worth mentioning that KSHV itself also upregulates PI3K/AKT signaling and inhibits cell apoptosis [115, 116]. A number of studies have found that different viral activators of PI3K/AKT pathway are used by Epstein-Barr virus (EBV) to infect the host cells latently and trigger carcinogenesis [107, 117, 118]. Inhibition of apoptosis and changes in cellular morphology are proposed to be the responsible measures taken by the virus PI3K/AKT pathway, which plays a role not only in coordination of apoptosis but also in cytoskeletal regulation of cell polarity and mobility. Viruses control penetration and intracellular transport of viral products via this pathway [107, 119]. Previous studies have identified viral activators of PI3K/AKT pathway in other viruses such as human papilloma virus type 16 (HPV16), paramyxoviruses like respiratory syncytial virus (RSV), poliovirus and rhinovirus, cardiovirus, encephalomyocarditis virus (EMCV), coxsackie virus, dengue virus, and Japanese encephalitis virus [120]. Some viruses such as influenza A virus, herpes simplex virus 1 (HSV-1), HIV, and HCV initiate the process as soon as entering the host cell. It is concluded that the early activation of PI3K/AKT pathway promotes viral entrance besides preparing the host environment for infection; for example, activated PI3K/AKT augments HCV entry and genome replication [121, 122]. In contrast, some viruses downregulate the activity of PI3K/AKI. Foot and mouth disease virus (FMDV) and measles virus are some examples of the aforementioned viruses. Some viral products of the latter facilitate PI3K/AKT activity. Interestingly, it has been shown that certain viral activators of PI3K/AKT also work as facilitators of cell apoptosis. A possible explanation for these blind spots is their function in different stages of viral life cycle. Over time, an extensive literature has been developed on the mechanisms of PI3K/AKT modulation by viruses; however, all the components are not fully understood [120]. Among coronaviruses, severe acute respiratory syndrome coronavirus (SARS CoV) has been shown to promote apoptosis by downregulating PI3K/AKT activity [123, 124]. At the same time, the virus has the ability to activate PI3K/AKT components by phosphorylation, prevent apoptosis, and establish a state of persistent infection in vitro [125, 126]. This is a phenomenon speculated to be responsible for prolonged presence of SARS-CoV particles in human feces in some patients. According to research, the activity level of PI3K/AKT pathway is not high enough to avoid apoptosis in a specific cell line [127, 128]. Meanwhile, some of the cells escaped apoptosis and were infected persistently [125].
A decade after SARS-CoV, a new coronavirus emerged and was named Middle East respiratory syndrome coronavirus (MERS-CoV) [129]. Kindrachuk and coworkers demonstrated the modulation of PI3K/AKT pathway during the infection and reduction in viral replication by adding PI3K/AKT inhibitors to host cells in vitro
[130]. Also, MERS-CoV promotes its replication by blocking autophagy [27, 131]. Furthermore, an indirect inhibitor of PI3K/AKT pathway, Saracatinib, has been identified to show antiviral effects against MERS-CoV [132]. Severe acute respiratory syndrome related coronavirus 2 (SARS-CoV-2) activates PI3K/AKT pathway as well [133-137].
Communication between the PI3K-Akt pathway and ACE2 activation in COVID-19 patients
The phosphatidylinositol 3-kinase (PI3K)/Akt pathway mediates different and various types of cell functions, including metabolism, growth, proliferation, and cell survival [138]. It was demonstrated that the virus enters the cell via two pathways; the non-endosomal pathway and endosomal pathway. In the non-endosomal entry, the virus genome is directly transmitted to the host cell’s cytosol through some molecules on the surface of the cell, including TMPRSS2 and furin. In the endosomal pathway, SARS-CoV-2 S protein can bind to CD147 and ACE2, letting the virus to enter the host cells and cause infection [93, 139]. In addition, it has been indicated that binding the virus spike proteins to furin and CD147 mediated with some signaling pathway, including PI3K/Akt signaling [140]. Also, it has been demonstrated that endocytosis of SARS-CoV-2 into the cell by receptor-mediated entry follows the clathrin-mediated pathway, regulated by PI3K/Akt signaling, which typically forms an early endosome and becomes a mature endosome before releasing contents in the cytosol [141]. Furthermore, it has been shown that the blocking of the PI3K/Akt signaling pathway has a suppression effect on the viruses that are utilizing clathrin-mediated pathway to enter the host cells [96]. ACE2 is expressed primarily by the lung vascular endothelial cells. However, it has been placed on the surface of some other extrapulmonary cells and tissues, such as the heart, nervous system, intestine, kidney, blood vessels, and muscle [142, 143]. The presence of this enzyme on the surface of various organs in the body may clarify the multiorgan failure and dysfunction in COVID-19 patients. As we know, the ACE2 acts as one of the SARS-CoV-2 receptors, and the reduction of ACE2 on the cells’ surface causes the improvement in producing angiotensin II (Ang II), which has been found in COVID-19 patients. Increasing the level of angiotensin II leads to increasing some inflammatory cytokines like interleukin (IL)-6 and TNF-α that consequently shows us the proinflammatory effect properties of Ang II. Increasing the level of Ang II stimulates fibrosis in different organs [97]. In addition, it is believed that angiotensin II (Ang II) type 1 receptor (AT1R) regulates most of the functions of the Ang II in the body. The AT1R exists in different organs and tissues, such as vascular smooth muscle, endothelium, heart, brain, kidney, adrenal gland, and adipose tissue, and assists most of the physiological functions stimulated by Ang II [144]. These different effects can be relevant to PI3K/AKT signaling pathway; by binding the Ang II to its receptor, AT1R, the PI3K/AKT signaling pathway is activated, and it may be responsible for those consequences [101] (Fig. 1).
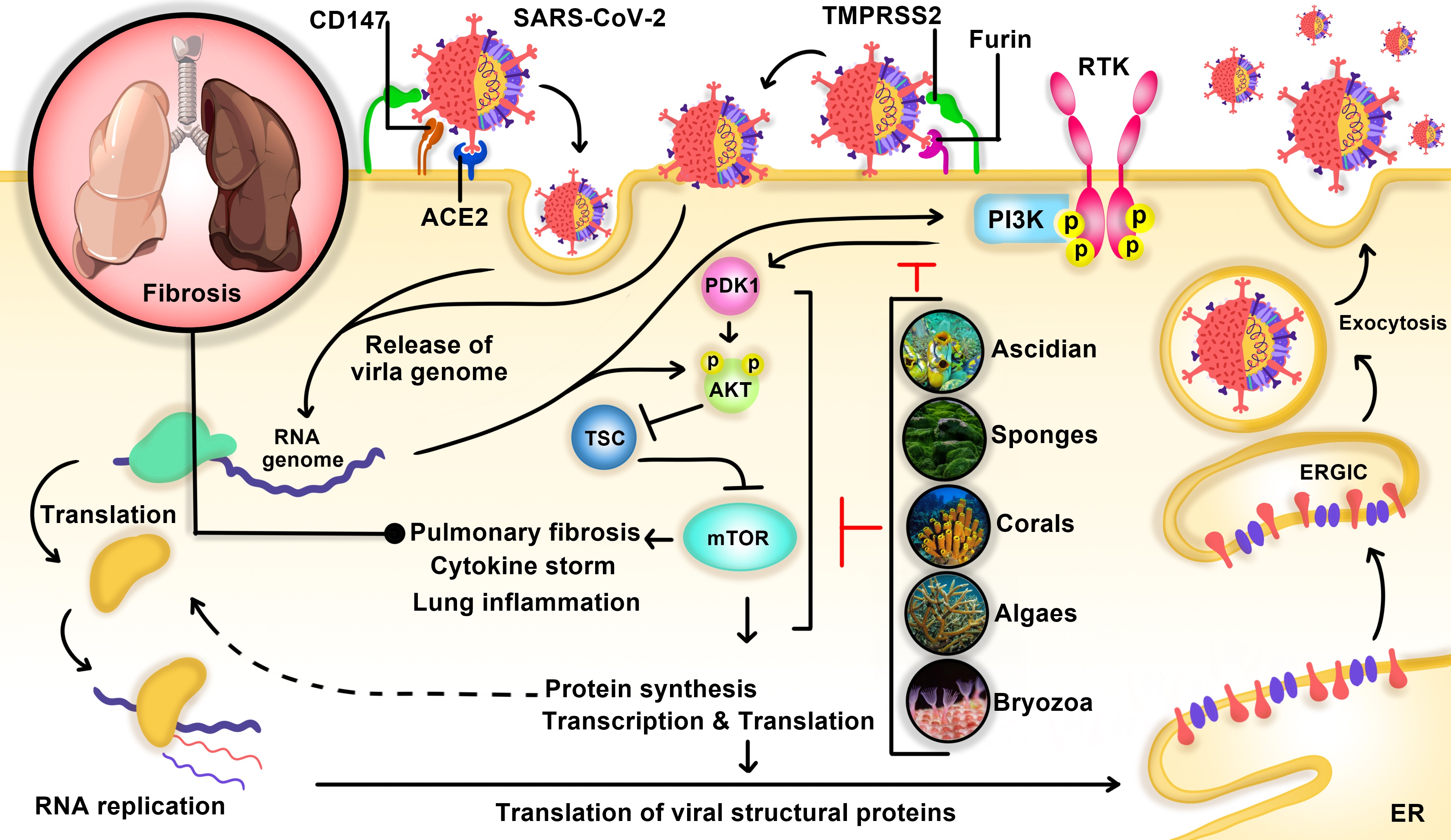
Fig. 1: The virus SARS-CoV-2 enters the cell via the non-endosomal pathway and/or endosomal pathway. In the Non-endosomal entry, the virus genome is directly transmitted to the host cell\'s cytosol through some molecules on the surface of the cell, including TMPRSS2 and furin. In the endosomal pathway, SARS-CoV-2 S protein can bind to angiotensin-converting enzyme 2 receptor (ACE2) and CD147 that let the virus enter the host cell and active PI3K/AKT/mTOR pathway, resulting in lung fibrosis. Ascidian, Sponges, Corals, Algaes, and Bryozoa are natural PI3K/AKT/mTOR inhibitors suppressing pulmonary fibrosis and cytokine storm.
Targeting PI3K/AKT in SARS-CoV-2: mechanisms
Since its emergence, SARS-CoV-2 has caused nearly 200 million diseased people and over 4 million deaths. Scientists are trying to find new therapeutic agents against the virus and lower the morbidity and mortality of the disease. SARS-CoV-2 has been shown to be an activator of ATR [136, 145], which serves as a part of PI3K/AKT pathway by detecting DNA damage, ceasing cellular proliferation, and upholding DNA integrity. It has been proposed that berzosertib acts as a protein kinase inhibitor of ATR and a potential antiviral agent against SARS-CoV-2. Berzosertib decreased viral production in vitro with minimal side effects compared with other protein kinase inhibitors. It also reduced infection associated programmed cell death [136, 146]. A strong relationship between PI3K/AKT activation and pulmonary fibrosis has been observed in SARS-CoV-2 induced pneumonia. D-Limonene an ingredient of citrus oil has been proposed as a potential antifibrotic agent in SARS-CoV-2 pulmonary fibrosis. D-Limonene not only suppresses PI3K/AKT expression but also inhibits its phosphorylation. Another possible explanation of D-Limonene induced inactivation of PI3K/AKT is reduction in reactive oxygen species (ROS) levels and TGF-β, which are known as the up-regulators of the pathway. Taken together D-Limonene improved chemical induced pulmonary fibrosis in rats and might be of use in SARS-CoV-2 induced pulmonary fibrosis, yet further research needs to be done [137]. Biguanides are other well-known PI3K/AKT inhibitors and are widely used as antidiabetic agents. An observational study showed lower incidence and morbidity of influenza among patients treated with biguanides compared with patients taking other antidiabetic agents. Another research on animals demonstrated lower pulmonary inflammation and mortality with better general condition in the biguanide treated group. Researchers assumed inhaled biguanides with lower systematic side effects may be useful for SARS-CoV-2 treatment [147]. It is worth mentioning that metformin, a biguanide antidiabetic medication-first used as anti-influenza drug has been demonstrated to be associated with lower mortality in COVID-19 patients However, there are some researches against its efficacy [148-150]. An increasing number of studies have found that dysregulated immune response and hyperinflammation are the reasons behind more severe SARS-CoV-2 infection and pulmonary fibrosis Mammalian target of rapamycin (mTOR), a kinase that functions in PI3K/AKT pathway and regulates inflammatory response by modulating different kinds of T cells and cytokine release. Inhibition of mTOR suppresses the cytokine storm, which maybe the cause of hyperinflammatory phase of COVID-19 [133, 151]. Inhibition of PI3K/AKT pathway by blocking mTOR has been claimed to be beneficial in SARS-CoV-2 infection [130, 152]. Sirolimus also known as rapamycin, an mTOR inhibitor used as an immunosuppressant against transplant rejection, was found to be advantageous in MERS-CoV. Its use might be beneficial in the hyper-inflammatory phase of SARS-CoV-2 infection [153-156]. Until 6 January 2021, 8 clinical trials have been investigating sirolimus use in COVID-19. Terrazzano and coworkers claimed everolimus, a rapamycin derivate, to be a potential antiviral agent against SARS-CoV-2 by blocking mTOR in PI3K/AKT pathway. Everolimus may decrease SARS-CoV-2 replication as it does in some other viral infections [152]. Azithromycin, a macrolide antibiotic has been used for its anti-inflammatory and immunomodulatory effects in some respiratory diseases. It inhibits PI3K/AKT pathway and thus reduces inflammatory cytokines level and modulates T cells. Along with that, Azithromycin may directly decrease viral entrance and replication via PI3K/AKT pathway modulation [157, 158] (Fig. 1, Table 1).
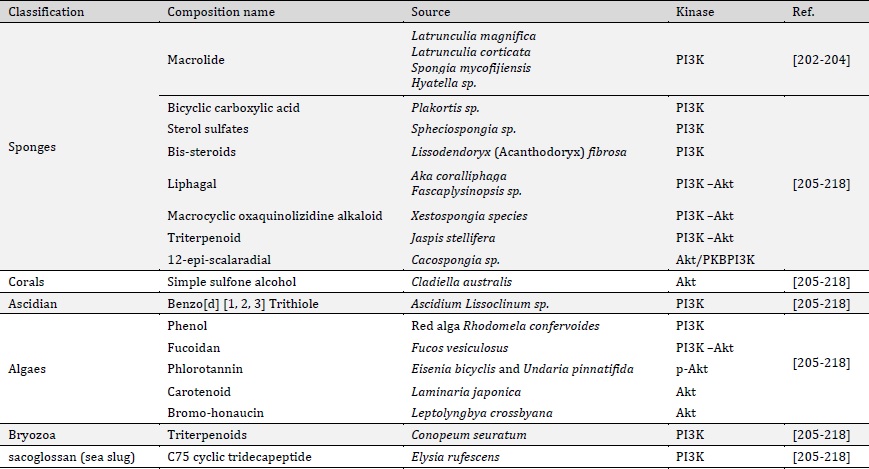
Table 1: Biologically active compounds from marine organisms capable of inhibiting PI3K/AKT
Sponges
Marine sponges (Phylum porifera) are some of the earliest multicellular invertebrate organisms. They are also a great source of valuable therapeutic natural products, some of which are thought to be important lead components for new therapy development. The majority of them are secondary compounds synthesized by sponges [159, 160], that might be used to protect them from harmful bacteria, algae, fungi, and other possible predators; a mechanism sponges have evolved over thousands of years of development. Marine sponges have a soft body, are immobile, and filter feeders, accumulating microscopic bits of food from the seawater that rises across their bodies. Bioactive compounds that come from sponges and other marine microorganisms have been found to contain anti-inflammatory, muscle relaxants, immunosuppressive, antiviral, antifungal, antibacterial anthelminthic, and antimalarial, properties. Sponge substances have exhibited a surprising range of chemical properties. In addition to rare nucleosides, marine sponges can synthesize other kinds of amino acid derivatives such as peroxides, alkaloids, sterols, cyclic peptides, fatty acids, terpenes, and so on. Antiviral compounds found in sponges are among the numerous innovative ways being used to put these compounds into medicinal applications [161-165].
A lot of molecules and anti-infective strategies have been developed for the protection of patients against virus and microbe attacks. These marine compounds can inhibit RNA and DNA viruses such as coronaviruses. They are in various structural classes such as peptides, terpenoids, alkaloids, polysaccharides, and steroids [165-168].
Macrolide
Macrolides are a class of antibiotics distinguished by the presence of a macrolide ring to which one or more deoxy sugars can be linked. It is common for lactone rings to have 14, 15, or 16 parts. Macrolides, which aggregate within leukocytes and are delivered to the site of infection, are used to treat Gram-positive bacterial infections of the respiratory and soft tissues, including S. pneumoniae and H. influenzae. Macrolides are known to inhibit the inflammatory process, minimize excessive production of cytokines in virus infection, and may lessen virus-related aggravation. Additionally, macrolides may affect phagocyte activity by altering a variety of processes such as chemotaxis, phagocytosis, oxidative burst, bacterial death, and cytokine production in addition to other effects [169-171] (Fig. 2).
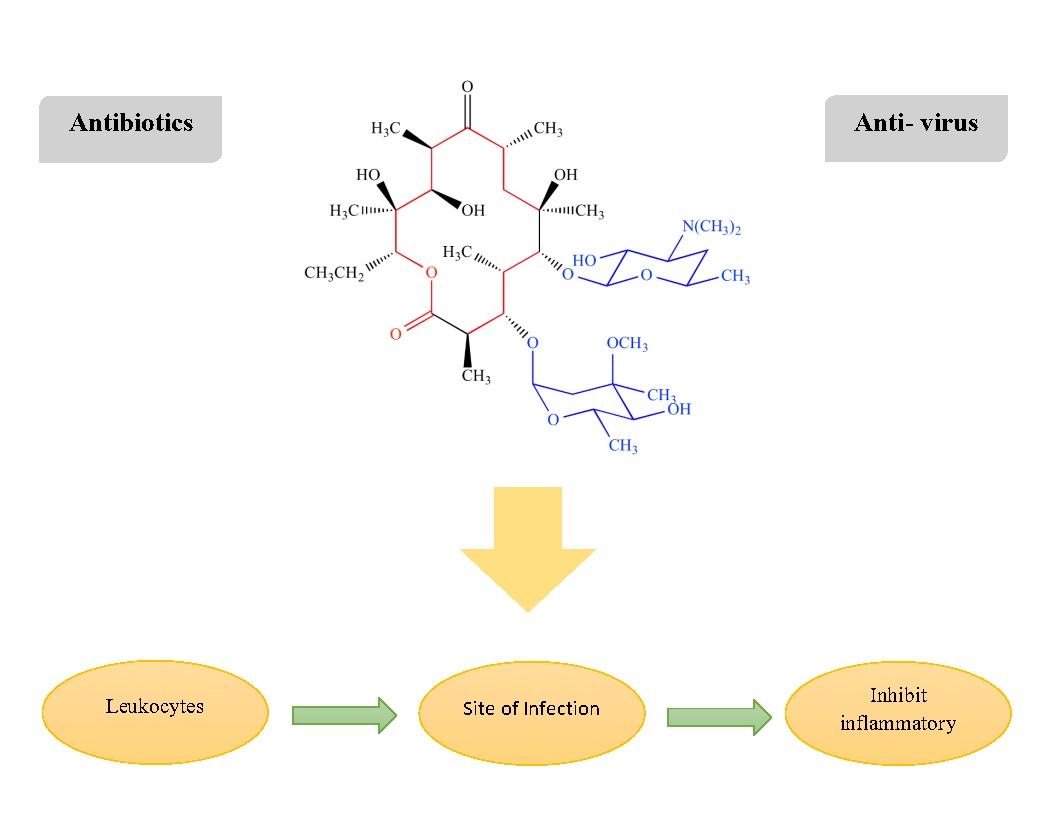
Fig. 2: Chemical structure: Macrolides
Fucoidan
Fucoidan is a sulfated anionic polysaccharide found in brown marine algae. A unique feature of this sulfated polysaccharide class is its fucose-rich skeleton, which is made up of a variety of sugars that vary considerably between species (e.g., mannose, galactose, glucose, xylose). Due to their low cytotoxicity and antiviral activity, sulfated polysaccharides have recently been shown to have antiviral activity both in vitro and in vivo. This raises the possibility of their usage in drug and gene delivery systems, wound healing formulations, and other applications. Fucoidans isolated from Adenocytis utricularis have inhibitory effects against the reproduction of a variety of enveloped viruses, such as the human immunodeficiency virus and the human cytomegalovirus. Activation of human neutrophils and natural killer cells (NK), as well as the generation of pro-inflammatory cytokines (IL-6, IL-8, and TNF-), were all enhanced by purified fucoidan from the brown seaweed Undaria pinnatifida, which also inhibited the cells’ spontaneous apoptosis. Dendritic cell maturation, cytotoxic T cell stimulation, Th1 immunological responses, antibody production in response to antigen exposure, and formation of memory T cells were all improved by fucoidan produced from Fucus vesiculosus [172] (Fig. 3, Fig. 4).
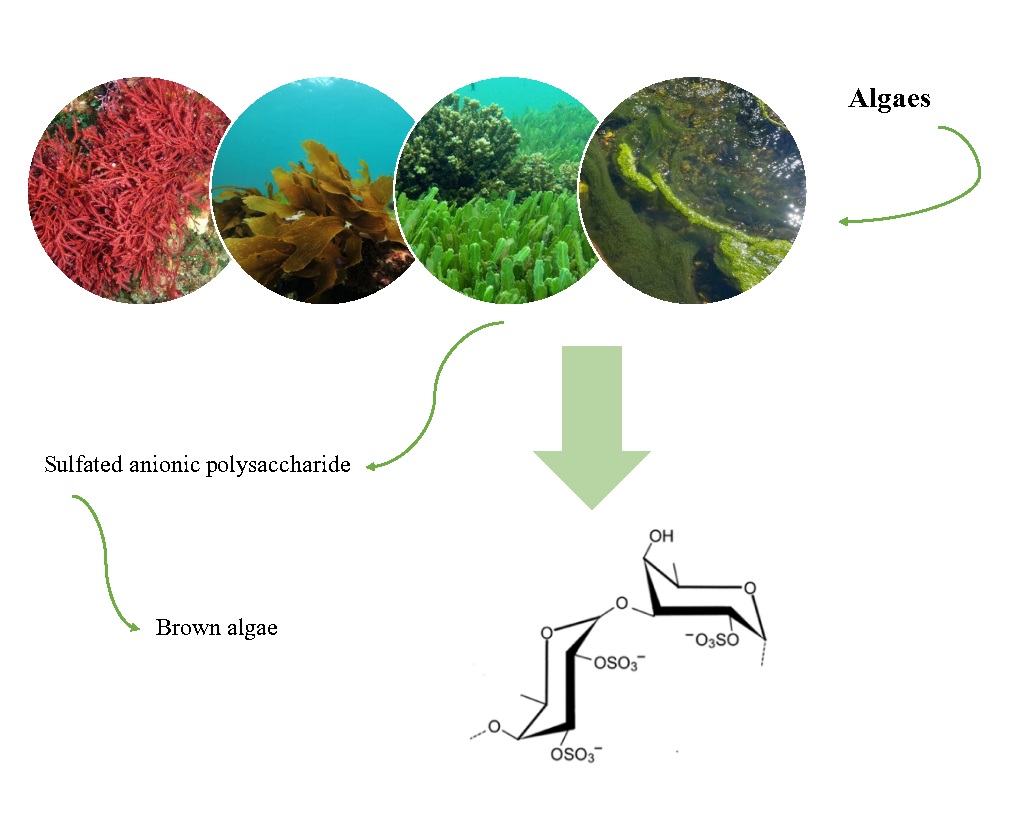
Fig. 3: Chemical structure: fucoidan (Fucos vesiculosus)
Fig. 4: Potential of fucoidan as a natural marine product that covers a wide range of antiviral activities, anti-bacterial, antioxidant, immunomodulatory, protective effect against oxidative stress and anti-inflammatory effects.
Association between idiopathic pulmonary fibrosis and COVID-19 severity: role of the PI3K/Akt route
Pulmonary fibrosis (PF) is initiated by a subgroup of macrophages named M2. M2 macrophages release transforming growth factor–β1 (TGF-β1), which activates fibroblasts and triggers fibrogenesis [173]. Furthermore, TGF-β1 activates PI3K/Akt pathway, by which autophagy and apoptosis of activated fibroblasts are suppressed and fibrotic remodeling is boosted [174-176]. SARS-CoV-2 may cause pneumonia followed by PF [177-180], which shares similar molecular basis with idiopathic pulmonary fibrosis (IPF) along with comparable role of PI3K/Akt pathway [181]. For instance, interleukin-6 (IL-6) and IL-8, proinflamatory cytokines upregulated by PI3K/Akt pathway [182-185], have been shown to be elevated in IPF and worsening its prognosis, with fibrogenic contribution to the disease [186, 187]. In the same way IL-6 and IL-8 are elevated during COVID-19 infection with significantly higher levels in severe disease [188-190]. Another example is significantly lower levels of Interferon-γ (IFN-γ) in patients with COVID-19 induced PF in comparison with COVID-19 patients without PF [191]. PI3k inhibitors enhance Interferon-γ (IFN-γ) levels in vivo [133]. Given the fact that SARS-CoV-2 is an activator of PI3K/Akt pathway [134-137, 192], lower level of IFN-γ in severe form of the disease (marked with PF [193] is understandable. PI3K/Akt cascades, specially some of its isoforms, are over expressed in IPF [194, 195]. Altogether, with the use of IFN-γ as a therapeutic agent in IPF [187-189], enhances our understanding of pathogenesis of both diseases. Moreover, positive genetic correlation between IPF and severe SARS-CoV-2 has been investigated [196-198]. Hence, there is no surprise that idiopathic pulmonary fibrosis (IPF) patients are more susceptible to get infected by COVID-19, with more severe disease and higher mortality rates [199-201].
Conclusion
The PI3K/AKT signaling pathway is known to be modulated by many viruses. Most viruses activate the pathway through different measures; and therefore, prevent cell apoptosis an effective and primitive host defense mechanism. Marine-derived natural products contain a variety of unique and fascinating structures that contribute significantly to biological activity and clinical therapeutic uses. Because of their bioactive and structure variety, natural compounds from marine organisms are regarded as an outstanding source of novel treatments with structural and chemical properties not present in the terrestrial environment. Several marine-derived kinase inhibitors have been developed from a variety of various sources and have been shown to inhibit a wide range of protein kinases. A huge spectrum of marine substances exhibits chemical structures with promising biological activity, indicating that they could be exploited to develop drugs for a variety of human diseases caused by viruses, notably COVID-19. Additionally, marine products have the benefit of being drug-like in nature with high levels of bioavailability, allowing them to be used as effective medications against viral illnesses in a very short period of time. Therefore, targeting the PI3K/Akt pathway can be useful as part of a COVID-19 response strategy. If this route is targeted early during the first phases of the disease, it has the potential to block the virus’s entry and replication and reduce the viral load, resulting in improved patient outcomes.Acknowledgements
Author Contributions
Koochaki P, Samami E, Ghoshouni H, Ghadimi D.J, Shadnoush F, Saberianpour Sh: Investigation; writing and preparation of the initial draft. Aleebrahim-Dehkordi E: Study design; Idea and the initiator of the figures; Project administration; writing-final draft and editing. Rafieian-Kopaei M: Supervised the project; editing and critically appraised the manuscript.
Disclosure Statement
The authors declare that no conflict of interests exists.
References
| 1 | Dhama K, Sharun K, Tiwari R, Dadar M, Malik YS, Singh KP, Chaicumpa W: COVID-19, an emerging coronavirus infection: advances and prospects in designing and developing vaccines, immunotherapeutics, and therapeutics. Hum Vaccin Immunother 2020;16:1232-1238. https://doi.org/10.1080/21645515.2020.1735227 |
| 2 | Huang J, Tao G, Liu J, Cai J, Huang Z, Chen JX: Current prevention of COVID-19: natural products and herbal medicine. Front Pharmacol 2020;11:588508. https://doi.org/10.3389/fphar.2020.588508 |
| 3 | Excler JL, Saville M, Berkley S, Kim JH: Vaccine development for emerging infectious diseases. Nat Med 2021;27:591-600. https://doi.org/10.1038/s41591-021-01301-0 |
| 4 | Berretta AA, Silveira MAD, Capcha JMC, De Jong D: Propolis and its potential against SARS-CoV-2 infection mechanisms and COVID-19 disease: Running title: Propolis against SARS-CoV-2 infection and COVID-19. Biomed Pharmacother 2020;131:110622. https://doi.org/10.1016/j.biopha.2020.110622 |
| 5 | Atanasov AG, Zotchev SB, Dirsch VM, Supuran CT: Natural products in drug discovery: advances and opportunities. Nat Rev Drug Discob 2021;20:200-216. https://doi.org/10.1038/s41573-020-00114-z |
| 6 | Omokhua-Uyi AG, Van Staden J: Natural product remedies for COVID-19: A focus on safety. S Afr J Bot 2021;139:386-398. https://doi.org/10.1016/j.sajb.2021.03.012 |
| 7 | Miller K, McGrath ME, Hu Z, Ariannejad S, Weston S, Frieman M, Jackson WT: Coronavirus interactions with the cellular autophagy machinery. Autophagy 2020;16:2131-2139. https://doi.org/10.1080/15548627.2020.1817280 |
| 8 | Yang N, Shen H-M: Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. Int J Biol Sci 2020;16:1724. https://doi.org/10.7150/ijbs.45498 |
| 9 | Racanelli AC, Kikkers SA, Choi AM, Cloonan SM: Autophagy and inflammation in chronic respiratory disease. Autophagy 2018;14:221-232. https://doi.org/10.1080/15548627.2017.1389823 |
| 10 | Brest P, Benzaquen J, Klionsky DJ, Hofman P, Mograbi B: Open questions for harnessing autophagy-modulating drugs in the SARS-CoV-2 war: hope or hype? Autophagy 2020;16:2267-2270. https://doi.org/10.1080/15548627.2020.1779531 |
| 11 | Pereira GJdS, Leão AHFF, Erustes AG, Morais IBdM, Vrechi TAdM, Zamarioli LdS, Pereira CAS, Marchioro LdO, Sperandio LP, Lins ÍVF: Pharmacological Modulators of Autophagy as a Potential Strategy for the Treatment of COVID-19. Int J Mol Sci 2021;22:4067. https://doi.org/10.3390/ijms22084067 |
| 12 | Schmidt U, Rein T: Novel treatment targets for COVID-19: Contribution from molecular psychiatry. World J Biol Psychiatry 2020;21:572-575. https://doi.org/10.1080/15622975.2020.1779344 |
| 13 | Zeng L, Li D, Tong W, Shi T, Ning B: Biochemical features and mutations of key proteins in SARS-CoV-2 and their impacts on RNA therapeutics. Biochem Pharmacol 2021;189:114424. https://doi.org/10.1016/j.bcp.2021.114424 |
| 14 | Sumon TA, Hussain MA, Hasan M, Rashid A, Abualreesh MH, Jang WJ, Sharifuzzaman S, Brown CL, Lee E-W, Hasan MT: Antiviral peptides from aquatic organisms: Functionality and potential inhibitory effect on SARS-CoV-2. Aquaculture 2021;541:736783. https://doi.org/10.1016/j.aquaculture.2021.736783 |
| 15 | Itani R, Tobaiqy M, Al Faraj A: Optimizing use of theranostic nanoparticles as a life-saving strategy for treating COVID-19 patients. Theranostics 2020;10:5932. https://doi.org/10.7150/thno.46691 |
| 16 | Zhou YW, Xie Y, Tang LS, Pu D, Zhu YJ, Liu JY, Ma XL: Therapeutic targets and interventional strategies in COVID-19: mechanisms and clinical studies. Signal Transduct Target Ther 2021;6:1-25. https://doi.org/10.1038/s41392-021-00733-x |
| 17 | Liu L, Zheng YY, Shao CL, Wang CY: Metabolites from marine invertebrates and their symbiotic microorganisms: Molecular diversity discovery, mining, and application. Mar Life Sci Technol 2019;1:60-94. https://doi.org/10.1007/s42995-019-00021-2 |
| 18 | Li T, Wang N, Zhang T, Zhang B, Sajeevan TP, Joseph V, Armstrong L, He S, Yan X, Naman CB: A systematic review of recently reported marine derived natural product kinase inhibitors. Mar Drugs 2019;17:493. https://doi.org/10.3390/md17090493 |
| 19 | Cragg GM, Newman DJ: Natural products: a continuing source of novel drug leads. Biochim Biophys Acta 2013;1830:3670-3695. https://doi.org/10.1016/j.bbagen.2013.02.008 |
| 20 | Molinski TF, Dalisay DS, Lievens SL, Saludes JP: Drug development from marine natural products. Nat Rev Drug Discov 2009;8:69-85. https://doi.org/10.1038/nrd2487 |
| 21 | Vanhaesebroeck B, Guillermet-Guibert J, Graupera M, Bilanges B: The emerging mechanisms of isoform-specific PI3K signalling. Nat Rev Mol Cell Biol 2010;11:329-341. https://doi.org/10.1038/nrm2882 |
| 22 | Yuan T, Cantley L: PI3K pathway alterations in cancer: variations on a theme. Oncogene 2008;27:5497-5510. https://doi.org/10.1038/onc.2008.245 |
| 23 | Khatpe AS, Adebayo AK, Herodotou CA, Kumar B, Nakshatri H: Nexus between PI3K/AKT and estrogen receptor signaling in breast cancer. Cancers (Basel) 2021;13:369. https://doi.org/10.3390/cancers13030369 |
| 24 | Engelman JA, Luo J, Cantley LC: The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet 2006;7:606-619. https://doi.org/10.1038/nrg1879 |
| 25 | Dienstmann R, Rodon J, Serra V, Tabernero J: Picking the Point of Inhibition: A Comparative Review of PI3K/AKT/mTOR Pathway InhibitorsDifferentiating PI3K/AKT/mTOR Pathway Inhibitors. Mol Cancer Ther 2014;13:1021-1031. https://doi.org/10.1158/1535-7163.MCT-13-0639 |
| 26 | Liu P, Cheng H, Roberts TM, Zhao JJ: Targeting the phosphoinositide 3-kinase pathway in cancer. Nat Rev Drug Discov 2009;8:627-644. https://doi.org/10.1038/nrd2926 |
| 27 | Yin Y, Dang W, Zhou X, Xu L, Wang W, Cao W, Chen S, Su J, Cai X, Xiao S: PI3K-Akt-mTOR axis sustains rotavirus infection via the 4E-BP1 mediated autophagy pathway and represents an antiviral target. Virulence 2018;9:83-98. https://doi.org/10.1080/21505594.2017.1326443 |
| 28 | Vanhaesebroeck B, Stephens L, Hawkins P: PI3K signalling: the path to discovery and understanding. Nat Rev Mol Cell Biol 2012;13:195-203. https://doi.org/10.1038/nrm3290 |
| 29 | Leverrier Y, Okkenhaug K, Sawyer C, Bilancio A, Vanhaesebroeck B, Ridley AJ: Class I phosphoinositide 3-kinase p110β is required for apoptotic cell and Fcγ receptor-mediated phagocytosis by macrophages. J Biol Chem 2003;278:38437-38442. https://doi.org/10.1074/jbc.M306649200 |
| 30 | Tamura N, Hazeki K, Okazaki N, Kametani Y, Murakami H, Takaba Y, Ishikawa Y, Nigorikawa K, Hazeki O: Specific role of phosphoinositide 3-kinase p110α in the regulation of phagocytosis and pinocytosis in macrophages. Biochem J 2009;423:99-108. https://doi.org/10.1042/BJ20090687 |
| 31 | Hawkins PT, Stephens LR, Suire S, Wilson M: PI3K signaling in neutrophils. Curr Top Microbiol Immunol 2010:183-202. https://doi.org/10.1007/82_2010_40 |
| 32 | Flannagan RS, Jaumouillé V, Grinstein S: The cell biology of phagocytosis. Annu Rev Pathol 2012;7:61-98. https://doi.org/10.1146/annurev-pathol-011811-132445 |
| 33 | Yoo SK, Deng Q, Cavnar PJ, Wu YI, Hahn KM, Huttenlocher A: Differential regulation of protrusion and polarity by PI (3) K during neutrophil motility in live zebrafish. Dev Cell 2010;18:226-236. https://doi.org/10.1016/j.devcel.2009.11.015 |
| 34 | Song MS, Salmena L, Pandolfi PP: The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 2012;13:283-296. https://doi.org/10.1038/nrm3330 |
| 35 | Patrucco E, Notte A, Barberis L, Selvetella G, Maffei A, Brancaccio M, Marengo S, Russo G, Azzolino O, Rybalkin SD: PI3Kγ modulates the cardiac response to chronic pressure overload by distinct kinase-dependent and-independent effects. Cell 2004;118:375-387. https://doi.org/10.1016/j.cell.2004.07.017 |
| 36 | Hirsch E, Braccini L, Ciraolo E, Morello F, Perino A: Twice upon a time: PI3K's secret double life exposed. Trends Biochem Sci 2009;34:244-248. https://doi.org/10.1016/j.tibs.2009.02.003 |
| 37 | Rauch J, Volinsky N, Romano D, Kolch W: The secret life of kinases: functions beyond catalysis. Cell Commun Signal 2011;9:1-28. https://doi.org/10.1186/1478-811X-9-23 |
| 38 | Falasca M, Maffucci T: Regulation and cellular functions of class II phosphoinositide 3-kinases. Biochem J 2012;443:587-601. https://doi.org/10.1042/BJ20120008 |
| 39 | Marat AL, Haucke V: Phosphatidylinositol 3‐phosphates-at the interface between cell signalling and membrane traffic. EMBO J 2016;35:561-579. https://doi.org/10.15252/embj.201593564 |
| 40 | Falasca M, Hughes WE, Dominguez V, Sala G, Fostira F, Fang MQ, Cazzolli R, Shepherd PR, James DE, Maffucci T: The role of phosphoinositide 3-kinase C2α in insulin signaling. J Biol Chem 2007;282:28226-28236. https://doi.org/10.1074/jbc.M704357200 |
| 41 | Wymann MP, Pirola L: Structure and function of phosphoinositide 3-kinases. Biochim Biophys 1998;1436:127-150. https://doi.org/10.1016/S0005-2760(98)00139-8 |
| 42 | Thoresen SB, Pedersen NM, Liestøl K, Stenmark H: A phosphatidylinositol 3-kinase class III sub-complex containing VPS15, VPS34, Beclin 1, UVRAG and BIF-1 regulates cytokinesis and degradative endocytic traffic. Exp Cell Res 2010;316:3368-3378. https://doi.org/10.1016/j.yexcr.2010.07.008 |
| 43 | Münz C: Macroautophagy-friend or foe of viral replication? EMBO Rep 2013;14:483-484. https://doi.org/10.1038/embor.2013.55 |
| 44 | Dong X, Levine B: Autophagy and viruses: adversaries or allies? J Innate Immun 2013;5:480-493. https://doi.org/10.1159/000346388 |
| 45 | Yuen CK, Wong WM, Mak LF, Wang X, Chu H, Yuen KY, Kok KH: Suppression of SARS‐CoV‐2 infection in ex‐vivo human lung tissues by targeting class III phosphoinositide 3‐kinase. J Med Virol 2021;93:2076-2083. https://doi.org/10.1002/jmv.26583 |
| 46 | Fruman DA, Meyers RE, Cantley LC: Phosphoinositide kinases. Annu Rev Biochem 1998;67:481. https://doi.org/10.1146/annurev.biochem.67.1.481 |
| 47 | Vanhaesebroeck B, Leevers SJ, Ahmadi K, Timms J, Katso R, Driscoll PC, Woscholski R, Parker PJ, Waterfield MD: Synthesis and function of 3-phosphorylated inositol lipids. Annu Rev Biochem 2001;70:535-602. https://doi.org/10.1146/annurev.biochem.70.1.535 |
| 48 | Katso R, Okkenhaug K, Ahmadi K, White S, Timms J, Waterfield MD: Cellular function of phosphoinositide 3-kinases: implications for development, immunity, homeostasis, and cancer. Annu Rev Cell Dev Biol 2001;17:615-675. https://doi.org/10.1146/annurev.cellbio.17.1.615 |
| 49 | Marshall A, Niiro H, Yun T, Clark E: Regulation of B-cell activation and differentiation by the phosphatidylinositol 3-kinase and phospholipase Cgamma pathway. Immunol Rev 2000;176:30-46. https://doi.org/10.1034/j.1600-065X.2000.00611.x |
| 50 | Kurosaki T, Okada T: Regulation of phospholipase C-γ2 and phosphoinositide 3-kinase pathways by adaptor proteins in B lymphocytes. Int Rev Immunol 2001;20:697-711. https://doi.org/10.3109/08830180109045586 |
| 51 | Turner H, Kinet J-P: Signalling through the high-affinity IgE receptor FcεRI. Nature 1999;402:24-30. https://doi.org/10.1038/35037021 |
| 52 | Jiang K, Zhong B, Gilvary DL, Corliss BC, Hong-Geller E, Wei S, Djeu JY: Pivotal role of phosphoinositide-3 kinase in regulation of cytotoxicity in natural killer cells. Nat Immunol 2000;1:419-425. https://doi.org/10.1038/80859 |
| 53 | Bone H, Williams NA: Antigen-receptor cross-linking and lipopolysaccharide trigger distinct phosphoinositide 3-kinase-dependent pathways to NF-κB activation in primary B cells. Int Immunol 2001;13:807-816. https://doi.org/10.1093/intimm/13.6.807 |
| 54 | Ishii KJ, Takeshita F, Gursel I, Gursel M, Conover J, Nussenzweig A, Klinman DM: Potential role of phosphatidylinositol 3 kinase, rather than DNA-dependent protein kinase, in CpG DNA-induced immune activation. J Exp Med 2002;196:269-274. https://doi.org/10.1084/jem.20020773 |
| 55 | Okada T, Maeda A, Iwamatsu A, Gotoh K, Kurosaki T: BCAP: the tyrosine kinase substrate that connects B cell receptor to phosphoinositide 3-kinase activation. Immunity 2000;13:817-827. https://doi.org/10.1016/S1074-7613(00)00079-0 |
| 56 | Chun Y, Kim J: Autophagy: an essential degradation program for cellular homeostasis and life. Cells 2018;7:278. https://doi.org/10.3390/cells7120278 |
| 57 | Kim J, Kundu M, Viollet B, Guan K-L: AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol 2011;13:132-141. https://doi.org/10.1038/ncb2152 |
| 58 | Echavarria-Consuegra L, Smit JM, Reggiori F: Role of autophagy during the replication and pathogenesis of common mosquito-borne flavi-and alphaviruses. Open Biol 2019;9:190009. https://doi.org/10.1098/rsob.190009 |
| 59 | Choi Y, Bowman JW, Jung JU: Autophagy during viral infection-a double-edged sword. Nat Rev Microbiol 2018;16:341-354. https://doi.org/10.1038/s41579-018-0003-6 |
| 60 | Jones RG, Elford AR, Parsons MJ, Wu L, Krawczyk CM, Yeh W-C, Hakem R, Rottapel R, Woodgett JR, Ohashi PS: CD28-dependent activation of protein kinase B/Akt blocks Fas-mediated apoptosis by preventing death-inducing signaling complex assembly. J Exp Med 2002;196:335-348. https://doi.org/10.1084/jem.20020307 |
| 61 | Liang Q, Luo Z, Zeng J, Chen W, Foo SS, Lee SA, Ge J, Wang S, Goldman SA, Zlokovic BV: Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 2016;19:663-671. https://doi.org/10.1016/j.stem.2016.07.019 |
| 62 | Cao B, Parnell LA, Diamond MS, Mysorekar IU: Inhibition of autophagy limits vertical transmission of Zika virus in pregnant mice. J Exp Med 2017;214:2303-2313. https://doi.org/10.1084/jem.20170957 |
| 63 | Zhou Y, Geng P, Liu Y, Wu J, Qiao H, Xie Y, Yin N, Chen L, Lin X, Liu Y: Rotavirus-encoded virus-like small RNA triggers autophagy by targeting IGF1R via the PI3K/Akt/mTOR pathway. Biochim Biophys Acta Mol Basis Dis 2018;1864:60-68. https://doi.org/10.1016/j.bbadis.2017.09.028 |
| 64 | Krejbich-Trotot P, Gay B, Li-Pat-Yuen G, Hoarau JJ, Jaffar-Bandjee MC, Briant L, Gasque P, Denizot M: Chikungunya triggers an autophagic process which promotes viral replication. Virol J 2011;8:1-10. https://doi.org/10.1186/1743-422X-8-432 |
| 65 | Wang R, Zhu Y, Zhao J, Ren C, Li P, Chen H, Jin M, Zhou H: Autophagy promotes replication of influenza A virus in vitro . J Virol 2019;93:e01984-01918. https://doi.org/10.1128/JVI.01984-18 |
| 66 | Bellacosa A, De Feo D, Godwin AK, Bell DW, Cheng JQ, Altomare DA, Wan M, Dubeau L, Scambia G, Masciullo V: Molecular alterations of the AKT2 oncogene in ovarian and breast carcinomas. Int J Cancer 1995;64:280-285. https://doi.org/10.1002/ijc.2910640412 |
| 67 | Bellacosa A, Kumar CC, Di Cristofano A, Testa JR: Activation of AKT kinases in cancer: implications for therapeutic targeting. Adv Cancer Res 2005;94:29-86. https://doi.org/10.1016/S0065-230X(05)94002-5 |
| 68 | Vara JÁF, Casado E, de Castro J, Cejas P, Belda-Iniesta C, González-Barón M: PI3K/Akt signalling pathway and cancer. Cancer Treat Rev 2004;30:193-204. https://doi.org/10.1016/j.ctrv.2003.07.007 |
| 69 | Zhao G-X, Pan H, Ouyang D-Y, He X-H: The critical molecular interconnections in regulating apoptosis and autophagy. Ann Med 2015;47:305-315. https://doi.org/10.3109/07853890.2015.1040831 |
| 70 | Duronio V: The life of a cell: apoptosis regulation by the PI3K/PKB pathway. Biochem J 2008;415:333-344. https://doi.org/10.1042/BJ20081056 |
| 71 | Song G, Ouyang G, Bao S: The activation of Akt/PKB signaling pathway and cell survival. J Cell Mol Med 2005;9:59-71. https://doi.org/10.1111/j.1582-4934.2005.tb00337.x |
| 72 | Coffer PJ, WOODGETT JR: Molecular cloning and characterisation of a novel putative protein‐serine kinase related to the cAMP‐dependent and protein kinase C families. Eur J Biochem 1991;201:475-481. https://doi.org/10.1111/j.1432-1033.1991.tb16305.x |
| 73 | Jones PF, Jakubowicz T, Pitossi FJ, Maurer F, Hemmings BA: Molecular cloning and identification of a serine/threonine protein kinase of the second-messenger subfamily. Proc Natl Acad Sci U S A 1991;88:4171-4175. https://doi.org/10.1073/pnas.88.10.4171 |
| 74 | Bellacosa A, Testa JR, Staal SP, Tsichlis PN: A retroviral oncogene, akt, encoding a serine-threonine kinase containing an SH2-like region. Science 1991;254:274-277. https://doi.org/10.1126/science.254.5029.274 |
| 75 | Burgering BMT, Coffer PJ: Protein kinase B (c-Akt) in phosphatidylinositol-3-OH kinase signal transduction. Nature 1995;376:599-602. https://doi.org/10.1038/376599a0 |
| 76 | Rameh LE, Cantley LC: The role of phosphoinositide 3-kinase lipid products in cell function. J Biol Chem 1999;274:8347-8350. https://doi.org/10.1074/jbc.274.13.8347 |
| 77 | Frech M, Andjelkovic M, Ingley E, Reddy KK, Falck JR, Hemmings BA: High affinity binding of inositol phosphates and phosphoinositides to the pleckstrin homology domain of RAC/protein kinase B and their influence on kinase activity. J Biol Chem 1997;272:8474-8481. https://doi.org/10.1074/jbc.272.13.8474 |
| 78 | Calleja V, Ameer-Beg SM, Vojnovic B, Woscholski R, Downward J, Larijani B: Monitoring conformational changes of proteins in cells by fluorescence lifetime imaging microscopy. Biochem J 2003;372:33-40. https://doi.org/10.1042/bj20030358 |
| 79 | Franke TF, Cantley LC: A Bad kinase makes good. Nature 1997;390:116-117. https://doi.org/10.1038/36442 |
| 80 | Meier R, Alessi DR, Cron P, Andjelkovic M, Hemmings BA: Mitogenic activation, phosphorylation, and nuclear translocation of protein kinase Bβ. J Biol Chem 1997;272:30491-30497. https://doi.org/10.1074/jbc.272.48.30491 |
| 81 | Andjelković M, Jakubowicz T, Cron P, Ming X-F, Han J-W, Hemmings BA: Activation and phosphorylation of a pleckstrin homology domain containing protein kinase (RAC-PK/PKB) promoted by serum and protein phosphatase inhibitors. Proc Natl Acad Sci U S A 1996;93:5699-5704. https://doi.org/10.1073/pnas.93.12.5699 |
| 82 | Zhou H, Summers SA, Birnbaum MJ, Pittman RN: Inhibition of Akt kinase by cell-permeable ceramide and its implications for ceramide-induced apoptosis. J Biol Chem 1998;273:16568-16575. https://doi.org/10.1074/jbc.273.26.16568 |
| 83 | Aman MJ, Lamkin TD, Okada H, Kurosaki T, Ravichandran KS: The inositol phosphatase SHIP inhibits Akt/PKB activation in B cells. J Biol Chem 1998;273:33922-33928. https://doi.org/10.1074/jbc.273.51.33922 |
| 84 | Astoul E, Watton S, Cantrell D: The dynamics of protein kinase B regulation during B cell antigen receptor engagement. J Cell Biol 1999;145:1511-1520. https://doi.org/10.1083/jcb.145.7.1511 |
| 85 | Guo H, German P, Bai S, Barnes S, Guo W, Qi X, Lou H, Liang J, Jonasch E, Mills GB: The PI3K/AKT pathway and renal cell carcinoma. J Genet Genomics 2015;42:343-353. https://doi.org/10.1016/j.jgg.2015.03.003 |
| 86 | Manning BD, Cantley LC: AKT/PKB signaling: navigating downstream. Cell 2007;129:1261-1274. https://doi.org/10.1016/j.cell.2007.06.009 |
| 87 | Hennessy BT, Smith DL, Ram PT, Lu Y, Mills GB: Exploiting the PI3K/AKT pathway for cancer drug discovery. Nat Rev Drug Discov 2005;4:988-1004. https://doi.org/10.1038/nrd1902 |
| 88 | Liu R, Chen Y, Liu G, Li C, Song Y, Cao Z, Li W, Hu J, Lu C, Liu Y: PI3K/AKT pathway as a key link modulates the multidrug resistance of cancers. Cell Death Dis 2020;11:1-12. https://doi.org/10.1038/s41419-020-02998-6 |
| 89 | Sun F, Mu C, Kwok HF, Xu J, Wu Y, Liu W, Sabatier JM, Annweiler C, Li X, Cao Z: Capivasertib restricts SARS-CoV-2 cellular entry: a potential clinical application for COVID-19. Int J Biol Sci 2021;17:2348. https://doi.org/10.7150/ijbs.57810 |
| 90 | Santamaria S: Targeting the PI3K/AKT pathway: a potential new weapon in the global fight against SARS-CoV-2? Int J Biol Sci 2021;17:2770. https://doi.org/10.7150/ijbs.63969 |
| 91 | Khezri MR: PI3K/AKT signaling pathway: a possible target for adjuvant therapy in COVID-19. Hum Cell 2021;34:700-701. https://doi.org/10.1007/s13577-021-00484-5 |
| 92 | Shah VK, Firmal P, Alam A, Ganguly D, Chattopadhyay S: Overview of immune response during SARS-CoV-2 infection: lessons from the past. Front Immunol 2020;11:1949. https://doi.org/10.3389/fimmu.2020.01949 |
| 93 | Lokhande AS, Devarajan PV: A review on possible mechanistic insights of Nitazoxanide for repurposing in COVID-19. Eur J Pharmacol 2021;891:173748. https://doi.org/10.1016/j.ejphar.2020.173748 |
| 94 | He Z, Khatib A-M, Creemers JW: Loss of proprotein convertase furin in mammary gland impairs proIGF1R and proIR processing and suppresses tumorigenesis in triple negative breast cancer. Cancers (Basel) 2020;12:2686. https://doi.org/10.3390/cancers12092686 |
| 95 | Lukiw WJ, Pogue A, Hill JM: SARS-CoV-2 infectivity and neurological targets in the brain. Cell Mol Neurobiol 2020:1-8. https://doi.org/10.1007/s10571-020-00947-7 |
| 96 | Cheng CY, Huang WR, Chi PI, Chiu HC, Liu HJ: Cell entry of bovine ephemeral fever virus requires activation of S rc‐JNK‐AP 1 and PI 3 K‐A kt‐NF‐κ B pathways as well as C ox‐2‐mediated PGE 2/EP receptor signalling to enhance clathrin‐mediated virus endocytosis. Cell Microbiol 2015;17:967-987. https://doi.org/10.1111/cmi.12414 |
| 97 | Miesbach W: Pathological role of angiotensin II in severe COVID-19. TH Open 2020;4:e138-e144. https://doi.org/10.1055/s-0040-1713678 |
| 98 | Gheblawi M, Wang K, Viveiros A, Nguyen Q, Zhong J-C, Turner AJ, Raizada MK, Grant MB, Oudit GY: Angiotensin-converting enzyme 2: SARS-CoV-2 receptor and regulator of the renin-angiotensin system: celebrating the 20th anniversary of the discovery of ACE2. Circ Res 2020;126:1456-1474. https://doi.org/10.1161/CIRCRESAHA.120.317015 |
| 99 | Aleebrahim-Dehkordi E, Reyhanian A, Hasanpour-Dehkordi A: Clinical Manifestation and the Risk of Exposure to SARS-CoV-2 (COVID‑19). Int J Prev Med 2020;11:86. |
| 100 | Ni W, Yang X, Yang D, Bao J, Li R, Xiao Y, Hou C, Wang H, Liu J, Yang D: Role of angiotensin-converting enzyme 2 (ACE2) in COVID-19. Crit Care 2020;24:1-10. https://doi.org/10.1186/s13054-020-03120-0 |
| 101 | Du N, Feng J, Hu LJ, Sun X, Sun HB, Zhao Y, Yang YP, Ren H: Angiotensin II receptor type 1 blockers suppress the cell proliferation effects of angiotensin II in breast cancer cells by inhibiting AT1R signaling. Oncol Rep 2012;27:1893-1903. |
| 102 | Zhang XL, Xing RG, Chen L, Liu CR, Miao ZG: PI3K/Akt signaling is involved in the pathogenesis of bleomycin‑induced pulmonary fibrosis via regulation of epithelial‑mesenchymal transition. Mol Med Report 2016;14:5699-5706. https://doi.org/10.3892/mmr.2016.5960 |
| 103 | Lee IT, Yang CM: Inflammatory signalings involved in airway and pulmonary diseases. Mediators Inflamm 2013;2013:791231. https://doi.org/10.1155/2013/791231 |
| 104 | Yodkeeree S, Ooppachai C, Pompimon W, Limtrakul P: O-methylbulbocapnine and dicentrine suppress LPS-induced inflammatory response by blocking NF-κB and AP-1 activation through inhibiting MAPKs and Akt signaling in RAW264.7 macrophages. Biol Pharm Bull 2018;41:1219-1227. https://doi.org/10.1248/bpb.b18-00037 |
| 105 | Wang J, Xie L, Wang S, Lin J, Liang J, Xu J: Azithromycin promotes alternatively activated macrophage phenotype in systematic lupus erythematosus via PI3K/Akt signaling pathway. Cell Death Dis 2018;9:1-13. https://doi.org/10.1038/s41419-018-1097-5 |
| 106 | Sun X, Chen L, He Z: PI3K/Akt-Nrf2 and anti-inflammation effect of macrolides in chronic obstructive pulmonary disease. Current drug metabolism 2019;20:301-304. https://doi.org/10.2174/1389200220666190227224748 |
| 107 | Ji WT, Liu HJ: PI3K-Akt signaling and viral infection. Recent Pat Biotechnol 2008;2:218-226. https://doi.org/10.2174/187220808786241042 |
| 108 | Teodoro JG, Branton PE: Regulation of apoptosis by viral gene products. J Virol 1997;71:1739-1746. https://doi.org/10.1128/jvi.71.3.1739-1746.1997 |
| 109 | Galluzzi L, Brenner C, Morselli E, Touat Z, Kroemer G: Viral control of mitochondrial apoptosis. PLoS Pathog 2008;4:e1000018. https://doi.org/10.1371/journal.ppat.1000018 |
| 110 | Rajala MS, Rajala RV, Astley RA, Butt AL, Chodosh J: Corneal cell survival in adenovirus type 19 infection requires phosphoinositide 3-kinase/Akt activation. J Virol 2005;79:12332-12341. https://doi.org/10.1128/JVI.79.19.12332-12341.2005 |
| 111 | Flaherty DM, Hinde SL, Monick MM, Powers LS, Bradford MA, Yarovinsky T, Hunninghake GW: Adenovirus vectors activate survival pathways in lung epithelial cells. Am J Physiol Lung Cell Physiol 2004;287:L393-L401. https://doi.org/10.1152/ajplung.00359.2003 |
| 112 | Xiang K, Wang B: Role of the PI3K‑AKT‑mTOR pathway in hepatitis B virus infection and replication. Mol Med Report 2018;17:4713-4719. https://doi.org/10.3892/mmr.2018.8395 |
| 113 | Guo H, Zhou T, Jiang D, Cuconati A, Xiao GH, Block TM, Guo JT: Regulation of hepatitis B virus replication by the phosphatidylinositol 3-kinase-akt signal transduction pathway. J Virol 2007;81:10072-10080. https://doi.org/10.1128/JVI.00541-07 |
| 114 | Zhou Q, Lui VW, Yeo W: Targeting the PI3K/Akt/mTOR pathway in hepatocellular carcinoma. Future oncology 2011;7:1149-1167. https://doi.org/10.2217/fon.11.95 |
| 115 | Deregibus MC, Cantaluppi V, Doublier S, Brizzi MF, Deambrosis I, Albini A, Camussi G: HIV-1-Tat protein activates phosphatidylinositol 3-kinase/AKT-dependent survival pathways in Kaposi's sarcoma cells. J Biol Chem 2002;277:25195-25202. https://doi.org/10.1074/jbc.M200921200 |
| 116 | Tomlinson CC, Damania B: The K1 protein of Kaposi's sarcoma-associated herpesvirus activates the Akt signaling pathway. J Virol 2004;78:1918-1927. https://doi.org/10.1128/JVI.78.4.1918-1927.2004 |
| 117 | Portis T, Longnecker R: Epstein-Barr virus (EBV) LMP2A mediates B-lymphocyte survival through constitutive activation of the Ras/PI3K/Akt pathway. Oncogene 2004;23:8619-8628. https://doi.org/10.1038/sj.onc.1207905 |
| 118 | Cooray S: The pivotal role of phosphatidylinositol 3-kinase-Akt signal transduction in virus survival. J Gen Virol 2004;85:1065-1076. https://doi.org/10.1099/vir.0.19771-0 |
| 119 | Dawson CW, Tramountanis G, Eliopoulos AG, Young LS: Epstein-Barr virus latent membrane protein 1 (LMP1) activates the phosphatidylinositol 3-kinase/Akt pathway to promote cell survival and induce actin filament remodeling. J Biol Chem 2003;278:3694-3704. https://doi.org/10.1074/jbc.M209840200 |
| 120 | Dunn EF, Connor JH: HijAkt: The PI3K/Akt pathway in virus replication and pathogenesis. Prog Mol Biol Transl Sci 2012;106:223-250. https://doi.org/10.1016/B978-0-12-396456-4.00002-X |
| 121 | Diehl N, Schaal H: Make yourself at home: viral hijacking of the PI3K/Akt signaling pathway. Viruses 2013;5:3192-3212. https://doi.org/10.3390/v5123192 |
| 122 | Shi Q, Hoffman B, Liu Q: PI3K-Akt signaling pathway upregulates hepatitis C virus RNA translation through the activation of SREBPs. Virology 2016;490:99-108. https://doi.org/10.1016/j.virol.2016.01.012 |
| 123 | Chan CM, Ma CW, Chan WY, Chan HYE: The SARS-coronavirus membrane protein induces apoptosis through modulating the Akt survival pathway. Arch Biochem Biophys 2007;459:197-207. https://doi.org/10.1016/j.abb.2007.01.012 |
| 124 | Surjit M, Liu B, Jameel S, Chow VT, Lal SK: The SARS coronavirus nucleocapsid protein induces actin reorganization and apoptosis in COS-1 cells in the absence of growth factors. Biochem J 2004;383:13-18. https://doi.org/10.1042/BJ20040984 |
| 125 | Mizutani T, Fukushi S, Saijo M, Kurane I, Morikawa S: JNK and PI3k/Akt signaling pathways are required for establishing persistent SARS-CoV infection in Vero E6 cells. Biochim Biophys Acta 2005;1741:4-10. https://doi.org/10.1016/j.bbadis.2005.04.004 |
| 126 | Mizutani T, Fukushi S, Ishii K, Sasaki Y, Kenri T, Saijo M, Kanaji Y, Shirota K, Kurane I, Morikawa S: Mechanisms of establishment of persistent SARS-CoV-infected cells. Biochem Biophys Res Commun 2006;347:261-265. https://doi.org/10.1016/j.bbrc.2006.06.086 |
| 127 | Chan PK, To KF, Lo AW, Cheung JL, Chu I, Au FW, Tong JH, Tam JS, Sung JJ, Ng HK: Persistent infection of SARS coronavirus in colonic cells in vitro . J Med Virol 2004;74:1-7. https://doi.org/10.1002/jmv.20138 |
| 128 | Mizutani T, Fukushi S, Saijo M, Kurane I, Morikawa S: Importance of Akt signaling pathway for apoptosis in SARS-CoV-infected Vero E6 cells. Virology 2004;327:169-174. https://doi.org/10.1016/j.virol.2004.07.005 |
| 129 | Dyall J, Gross R, Kindrachuk J, Johnson RF, Olinger GG, Hensley LE, Frieman MB, Jahrling PB: Middle East respiratory syndrome and severe acute respiratory syndrome: current therapeutic options and potential targets for novel therapies. Drugs 2017;77:1935-1966. https://doi.org/10.1007/s40265-017-0830-1 |
| 130 | Kindrachuk J, Ork B, Mazur S, Holbrook MR, Frieman MB, Traynor D, Johnson RF, Dyall J, Kuhn JH, Olinger GG: Antiviral potential of ERK/MAPK and PI3K/AKT/mTOR signaling modulation for Middle East respiratory syndrome coronavirus infection as identified by temporal kinome analysis. Antimicrob Agents Chemother 2015;59:1088-1099. https://doi.org/10.1128/AAC.03659-14 |
| 131 | Gassen NC, Niemeyer D, Muth D, Corman VM, Martinelli S, Gassen A, Hafner K, Papies J, Mösbauer K, Zellner A: SKP2 attenuates autophagy through Beclin1-ubiquitination and its inhibition reduces MERS-Coronavirus infection. Nat Commun 2019;10:1-16. https://doi.org/10.1038/s41467-019-13659-4 |
| 132 | Shin JS, Jung E, Kim M, Baric RS, Go YY: Saracatinib inhibits middle east respiratory syndrome-coronavirus replication in vitro . Viruses 2018;10:283. https://doi.org/10.3390/v10060283 |
| 133 | Terrazzano G, Rubino V, Palatucci AT, Giovazzino A, Carriero F, Ruggiero G: An open question: is it rational to inhibit the mTor-dependent pathway as COVID-19 therapy? Front Pharmacol 2020;11:856. https://doi.org/10.3389/fphar.2020.00856 |
| 134 | Bouhaddou M, Memon D, Meyer B, White KM, Rezelj VV, Marrero MC, Polacco BJ, Melnyk JE, Ulferts S, Kaake RM: The global phosphorylation landscape of SARS-CoV-2 infection. Cell 2020;182:685-712.e19. https://doi.org/10.1016/j.cell.2020.06.034 |
| 135 | Appelberg S, Gupta S, Svensson Akusjärvi S, Ambikan AT, Mikaeloff F, Saccon E, Végvári Á, Benfeitas R, Sperk M, Ståhlberg M: Dysregulation in Akt/mTOR/HIF-1 signaling identified by proteo-transcriptomics of SARS-CoV-2 infected cells. Emerg Microbes Infect 2020;9:1748-1760. https://doi.org/10.1080/22221751.2020.1799723 |
| 136 | Garcia Jr G, Sharma A, Ramaiah A, Sen C, Purkayastha A, Kohn DB, Parcells MS, Beck S, Kim H, Bakowski MA: Antiviral drug screen identifies DNA-damage response inhibitor as potent blocker of SARS-CoV-2 replication. Cell Rep 2021;35:108940. https://doi.org/10.1016/j.celrep.2021.108940 |
| 137 | Yang F, Chen R, Li WY, Zhu HY, Chen XX, Hou ZF, Cao RS, Zang G, Li YX, Zhang W: D-limonene is a potential monoterpene to inhibit PI3K/Akt/IKK-α/NF-κB p65 signaling pathway in coronavirus disease 2019 pulmonary fibrosis. Front Med (Lausanne) 2021;8:591830. https://doi.org/10.3389/fmed.2021.591830 |
| 138 | Vivanco I, Sawyers CL: The phosphatidylinositol 3-kinase-AKT pathway in human cancer. Nat Rev Cancer 2002;2:489-501. https://doi.org/10.1038/nrc839 |
| 139 | Sharifkashani S, Bafrani MA, Khaboushan AS, Pirzadeh M, Kheirandish A, Yavarpour_Bali H, Hessami A, Saghazadeh A, Rezaei N: Angiotensin-converting enzyme 2 (ACE2) receptor and SARS-CoV-2: potential therapeutic targeting. Eur J Pharmacol 2020;884:173455. https://doi.org/10.1016/j.ejphar.2020.173455 |
| 140 | Shou J, Wang M, Cheng X, Wang X, Zhang L, Liu Y, Fei C, Wang C, Gu F, Xue F: Tizoxanide induces autophagy by inhibiting PI3K/Akt/mTOR pathway in RAW264 7 macrophage cells. Arch Pharm Res 2020;43:257-270. https://doi.org/10.1007/s12272-019-01202-4 |
| 141 | Glebov OO: Understanding SARS‐CoV‐2 endocytosis for COVID‐19 drug repurposing. FEBS J 2020;287:3664-3671. https://doi.org/10.1111/febs.15369 |
| 142 | Forrester SJ, Booz GW, Sigmund CD, Coffman TM, Kawai T, Rizzo V, Scalia R, Eguchi S: Angiotensin II signal transduction: an update on mechanisms of physiology and pathophysiology. Physiol Rev 2018;98:1627-1738. https://doi.org/10.1152/physrev.00038.2017 |
| 143 | Hamming I, Timens W, Bulthuis M, Lely A, Navis Gv, van Goor H: Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J Pathol 2004;203:631-637. https://doi.org/10.1002/path.1570 |
| 144 | De Gasparo M, Catt K, Inagami T, Wright J, Unger T: International union of pharmacology. XXIII. The angiotensin II receptors. Pharmacol Rev 2000;52:415-472. |
| 145 | Nicola M, Alsafi Z, Sohrabi C, Kerwan A, Al-Jabir A, Iosifidis C, Agha M, Agha R: The socio-economic implications of the coronavirus pandemic (COVID-19): A review. Int J Surg 2020;78:185-193. https://doi.org/10.1016/j.ijsu.2020.04.018 |
| 146 | Gorecki L, Andrs M, Rezacova M, Korabecny J: Discovery of ATR kinase inhibitor berzosertib (VX-970, M6620): Clinical candidate for cancer therapy. Pharmacol Ther 2020;210:107518. https://doi.org/10.1016/j.pharmthera.2020.107518 |
| 147 | Lehrer S: Inhaled biguanides and mTOR inhibition for influenza and coronavirus. World Acad Sci J 2020;2:1-1. https://doi.org/10.3892/wasj.2020.42 |
| 148 | Karam BS, Morris RS, Bramante CT, Puskarich M, Zolfaghari EJ, Lotfi‐Emran S, Ingraham NE, Charles A, Odde DJ, Tignanelli CJ: mTOR inhibition in COVID‐19: A commentary and review of efficacy in RNA viruses. J Med Virol 2021;93:1843-1846. https://doi.org/10.1002/jmv.26728 |
| 149 | Luo P, Qiu L, Liu Y, Liu Xl, Zheng JL, Xue HY, Liu WH, Liu D, Li J: Metformin treatment was associated with decreased mortality in COVID-19 patients with diabetes in a retrospective analysis. Am J Trop Med Hyg 2020;103:69-72. https://doi.org/10.4269/ajtmh.20-0375 |
| 150 | Cariou B, Hadjadj S, Wargny M, Pichelin M, Al-Salameh A, Allix I, Amadou C, Arnault G, Baudoux F, Bauduceau B: Phenotypic characteristics and prognosis of inpatients with COVID-19 and diabetes: the CORONADO study. Diabetologia 2020;63:1500-1515. https://doi.org/10.1007/s00125-020-05180-x |
| 151 | Felsenstein S, Herbert JA, McNamara PS, Hedrich CM: COVID-19: Immunology and treatment options. Clin Immunol 2020;215:108448. https://doi.org/10.1016/j.clim.2020.108448 |
| 152 | Romanelli A, Mascolo S: Sirolimus to treat SARS-CoV-2 infection: an old drug for a new disease. Respir Med 2020;8:420-422. https://doi.org/10.34172/jrcm.2020.044 |
| 153 | Husain A, Byrareddy SN: Rapamycin as a potential repurpose drug candidate for the treatment of COVID-19 Chem Biol Interact 2020;331:109282. https://doi.org/10.1016/j.cbi.2020.109282 |
| 154 | Patocka J, Kuca K, Oleksak P, Nepovimova E, Valis M, Novotny M, Klimova B: Rapamycin: drug repurposing in SARS-CoV-2 infection. Pharmaceuticals 2021;14:217. https://doi.org/10.3390/ph14030217 |
| 155 | Bischof E, Siow RC, Zhavoronkov A, Kaeberlein M: The potential of rapalogs to enhance resilience against SARS-CoV-2 infection and reduce the severity of COVID-19. Lancet Healthy Longev 2021;2:e105-e111. https://doi.org/10.1016/S2666-7568(20)30068-4 |
| 156 | Pani A, Lauriola M, Romandini A, Scaglione F: Macrolides and viral infections: focus on azithromycin in COVID-19 pathology. Int J Antimicrob Agents 2020;56:106053. https://doi.org/10.1016/j.ijantimicag.2020.106053 |
| 157 | Khezri MR, Zolbanin NM, Ghasemnejad-Berenji M, Jafari R: Azithromycin: Immunomodulatory and antiviral properties for SARS-CoV-2 infection. Eur J Pharmacol 2021;905:174191. https://doi.org/10.1016/j.ejphar.2021.174191 |
| 158 | Saito A, Horie M, Nagase T: TGF-β signaling in lung health and disease. Int J Mol Sci 2018;19:2460. https://doi.org/10.3390/ijms19082460 |
| 159 | Sarma AS, Daum T, Müller WE: Secondary metabolites from marine sponges. Ullstein Mosby, Berlin, 1993. |
| 160 | Sagar S, Kaur M, Minneman KP: Antiviral lead compounds from marine sponges. Mar Drugs 2010;8:2619-2638. https://doi.org/10.3390/md8102619 |
| 161 | Hadas E, Shpigel M, Ilan M: Particulate organic matter as a food source for a coral reef sponge. J Exp Biol 2009;212:3643-3650. https://doi.org/10.1242/jeb.027953 |
| 162 | Donia M, Hamann MT: Marine natural products and their potential applications as anti-infective agents. Lancet Infect Dis 2003;3:338-348. https://doi.org/10.1016/S1473-3099(03)00655-8 |
| 163 | Blunt JW, Copp BR, Keyzers RA, Munro MH, Prinsep MR: Marine natural products. Nat Prod Rep 2013;30:237-323. https://doi.org/10.1039/C2NP20112G |
| 164 | Anjum K, Abbas SQ, Shah SAA, Akhter N, Batool S: Marine sponges as a drug treasure. Biomol Ther (Seoul) 2016;24:347-362. https://doi.org/10.4062/biomolther.2016.067 |
| 165 | Zaporozhets TS, Besednova NN: Biologically active compounds from marine organisms in the strategies for combating coronaviruses. AIMS Microbiol 2020;6:470. https://doi.org/10.3934/microbiol.2020028 |
| 166 | Malve H: Exploring the ocean for new drug developments: Marine pharmacology. J Pharm Bioallied Sci 2016;8:83. https://doi.org/10.4103/0975-7406.171700 |
| 167 | Cheung RCF, Wong JH, Pan W, Chan YS, Yin C, Dan X, Ng TB: Marine lectins and their medicinal applications. Appl Microbiol Biotechnol 2015;99:3755-3773. https://doi.org/10.1007/s00253-015-6518-0 |
| 168 | Gentile D, Patamia V, Scala A, Sciortino MT, Piperno A, Rescifina A: Putative inhibitors of SARS-CoV-2 main protease from a library of marine natural products: a virtual screening and molecular modeling study. Mar Drugs 2020;18:225. https://doi.org/10.3390/md18040225 |
| 169 | Min JY, Jang YJ: Macrolide therapy in respiratory viral infections. Mediators Inflamm 2012;2012:649570. https://doi.org/10.1155/2012/649570 |
| 170 | Buret AG: Immuno-modulation and anti-inflammatory benefits of antibiotics: the example of tilmicosin. Can J Vet Res 2010;74:1-10. |
| 171 | Kanoh S, Rubin BK: Mechanisms of action and clinical application of macrolides as immunomodulatory medications. Clin Microbiol Rev 2010;23:590-615. https://doi.org/10.1128/CMR.00078-09 |
| 172 | Jin JO, Zhang W, Du JY, Wong KW, Oda T, Yu Q: Fucoidan can function as an adjuvant in vivo to enhance dendritic cell maturation and function and promote antigen-specific T cell immune responses. PLoS One 2014;9:e99396. https://doi.org/10.1371/journal.pone.0099396 |
| 173 | Horowitz JC, Rogers DS, Sharma V, Vittal R, White ES, Cui Z, Thannickal VJ: Combinatorial activation of FAK and AKT by transforming growth factor-β1 confers an anoikis-resistant phenotype to myofibroblasts. Cell Signal 2007;19:761-771. https://doi.org/10.1016/j.cellsig.2006.10.001 |
| 174 | Larson-Casey JL, Deshane JS, Ryan AJ, Thannickal VJ, Carter AB: Macrophage Akt1 kinase-mediated mitophagy modulates apoptosis resistance and pulmonary fibrosis. Immunity 2016;44:582-596. https://doi.org/10.1016/j.immuni.2016.01.001 |
| 175 | Xia H, Diebold D, Nho R, Perlman D, Kleidon J, Kahm J, Avdulov S, Peterson M, Nerva J, Bitterman P: Pathological integrin signaling enhances proliferation of primary lung fibroblasts from patients with idiopathic pulmonary fibrosis. J Exp Med 2008;205:1659-1672. https://doi.org/10.1084/jem.20080001 |
| 176 | González J, Benítez ID, Carmona P, Santisteve S, Monge A, Moncusí-Moix A, Gort-Paniello C, Pinilla L, Carratalá A, Zuil M: Pulmonary function and radiologic features in survivors of critical COVID-19: a 3-month prospective cohort. Chest 2021;160:187-198. |
| 177 | Yao XH, Luo T, Shi Y, He ZC, Tang R, Zhang PP, Cai J, Zhou XD, Jiang DP, Fei XC: A cohort autopsy study defines COVID-19 systemic pathogenesis. Cell Res 2021;31:836-846. https://doi.org/10.1038/s41422-021-00523-8 |
| 178 | Zhao YM, Shang YM, Song WB, Li QQ, Xie H, Xu QF, Jia JL, Li LM, Mao HL, Zhou XM: Follow-up study of the pulmonary function and related physiological characteristics of COVID-19 survivors three months after recovery. EClinicalMedicine 2020;25:100463. https://doi.org/10.1016/j.eclinm.2020.100463 |
| 179 | Shi H, Han X, Jiang N, Cao Y, Alwalid O, Gu J, Fan Y, Zheng C: Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: a descriptive study. Lancet Infect Dis 2020;20:425-434. https://doi.org/10.1016/S1473-3099(20)30086-4 |
| 180 | Wang Y, Zhang L, Wu GR, Zhou Q, Yue H, Rao LZ, Yuan T, Mo B, Wang FX, Chen LM: MBD2 serves as a viable target against pulmonary fibrosis by inhibiting macrophage M2 program. Sci Adv 2021;7:eabb6075. https://doi.org/10.1126/sciadv.abb6075 |
| 181 | Zegeye MM, Lindkvist M, Fälker K, Kumawat AK, Paramel G, Grenegård M, Sirsjö A, Ljungberg LU: Activation of the JAK/STAT3 and PI3K/AKT pathways are crucial for IL-6 trans-signaling-mediated pro-inflammatory response in human vascular endothelial cells. Cell Commun Signal 2018;16:1-10. https://doi.org/10.1186/s12964-018-0268-4 |
| 182 | Tong KM, Shieh DC, Chen CP, Tzeng CY, Wang SP, Huang KC, Chiu YC, Fong YC, Tang CH: Leptin induces IL-8 expression via leptin receptor, IRS-1, PI3K, Akt cascade and promotion of NF-κB/p300 binding in human synovial fibroblasts. Cell Signal 2008;20:1478-1488. https://doi.org/10.1016/j.cellsig.2008.04.003 |
| 183 | Wang L, Tang C, Cao H, Li K, Pang X, Zhong L, Dang W, Tang H, Huang Y, Wei L: Activation of IL-8 via PI3K/Akt-dependent pathway is involved in leptin-mediated epithelial-mesenchymal transition in human breast cancer cells. Cancer Biol Ther 2015;16:1220-1230. https://doi.org/10.1080/15384047.2015.1056409 |
| 184 | Bickel M: The role of interleukin-8 in inflammation and mechanisms of regulation. J Periodontol 1993;64:456-460. |
| 185 | Papiris SA, Tomos IP, Karakatsani A, Spathis A, Korbila I, Analitis A, Kolilekas L, Kagouridis K, Loukides S, Karakitsos P: High levels of IL-6 and IL-8 characterize early-on idiopathic pulmonary fibrosis acute exacerbations. Cytokine 2018;102:168-172. https://doi.org/10.1016/j.cyto.2017.08.019 |
| 186 | Yang L, Herrera J, Gilbertsen A, Xia H, Smith K, Benyumov A, Bitterman PB, Henke CA: IL-8 mediates idiopathic pulmonary fibrosis mesenchymal progenitor cell fibrogenicityAm J Physiol Cell Mol Physiol 2018;314:L127-L136. https://doi.org/10.1152/ajplung.00200.2017 |
| 187 | Del Valle DM, Kim-Schulze S, Huang H-H, Beckmann ND, Nirenberg S, Wang B, Lavin Y, Swartz TH, Madduri D, Stock A: An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med 2020;26:1636-1643. https://doi.org/10.1038/s41591-020-1051-9 |
| 188 | Gong J, Dong H, Xia QS, Huang ZY, Wang DK, Zhao Y, Liu WH, Tu SH, Zhang MM, Wang Q: Correlation analysis between disease severity and inflammation-related parameters in patients with COVID-19: a retrospective study. BMC Infect Dis 2020;20:1-7. https://doi.org/10.1186/s12879-020-05681-5 |
| 189 | Nagant C, Ponthieux F, Smet J, Dauby N, Doyen V, Besse-Hammer T, De Bels D, Maillart E, Corazza F: A score combining early detection of cytokines accurately predicts COVID-19 severity and intensive care unit transfer. Int J Infect Dis 2020;101:342-345. https://doi.org/10.1016/j.ijid.2020.10.003 |
| 190 | Hu ZJ, Xu J, Yin JM, Li L, Hou W, Zhang LL, Zhou Z, Yu YZ, Li HJ, Feng YM: Lower circulating interferon-gamma is a risk factor for lung fibrosis in COVID-19 patients. Front Immunol 2020;11:585647. https://doi.org/10.3389/fimmu.2020.585647 |
| 191 | Marshall NA, Galvin KC, Corcoran A-MB, Boon L, Higgs R, Mills KH: Immunotherapy with PI3K Inhibitor and Toll-Like Receptor Agonist Induces IFN-γ+ IL-17+ Polyfunctional T Cells That Mediate Rejection of Murine TumorsPI3K Inhibitors Improve Efficacy of TLR-Based Tumor Vaccine. Cancer Res 2012;72:581-591. https://doi.org/10.1158/0008-5472.CAN-11-0307 |
| 192 | Garcia-Revilla J, Deierborg T, Venero JL, Boza-Serrano A: Hyperinflammation and fibrosis in severe COVID-19 patients: galectin-3, a target molecule to consider. Front Immunol 2020;11:2069. https://doi.org/10.3389/fimmu.2020.02069 |
| 193 | Conte E, Fruciano M, Fagone E, Gili E, Caraci F, Iemmolo M, Crimi N, Vancheri C: Inhibition of PI3K prevents the proliferation and differentiation of human lung fibroblasts into myofibroblasts: the role of class I P110 isoforms. PLoS One 2011;6:e24663. https://doi.org/10.1371/journal.pone.0024663 |
| 194 | Conte E, Gili E, Fruciano M, Korfei M, Fagone E, Iemmolo M, Lo Furno D, Giuffrida R, Crimi N, Guenther A: PI3K p110γ overexpression in idiopathic pulmonary fibrosis lung tissue and fibroblast cells: in vitro effects of its inhibition. Lab Invest 2013;93:566-576. https://doi.org/10.1038/labinvest.2013.6 |
| 195 | Staab-Weijnitz CA, Eickelberg O: Repositioning compounds from cancer drug discovery to IPF: PI3K inhibition. Thorax 2016;71:675-676. https://doi.org/10.1136/thoraxjnl-2016-208680 |
| 196 | Lukey PT, Harrison SA, Yang S, Man Y, Holman BF, Rashidnasab A, Azzopardi G, Grayer M, Simpson JK, Bareille P: A randomised, placebo-controlled study of omipalisib (PI3K/mTOR) in idiopathic pulmonary fibrosis. Eur Respir J 2019;53:1801992. https://doi.org/10.1183/13993003.01992-2018 |
| 197 | Lin S, Jin J, Liu Y, Tian H, Zhang Y, Fu R, Zhang J, Wang M, Du T, Ji M: Discovery of 4-methylquinazoline based PI3K inhibitors for the potential treatment of idiopathic pulmonary fibrosis. J Med Chem 2019;62:8873-8879. https://doi.org/10.1021/acs.jmedchem.9b00969 |
| 198 | Wang L, Balmat TJ, Antonia AL, Constantine FJ, Henao R, Burke TW, Ingham A, McClain MT, Tsalik EL, Ko ER: An atlas connecting shared genetic architecture of human diseases and molecular phenotypes provides insight into COVID-19 susceptibility. Genome Med 2021;13:1-19. https://doi.org/10.1186/s13073-021-00904-z |
| 199 | Taz TA, Ahmed K, Paul BK, Kawsar M, Aktar N, Mahmud SH, Moni MA: Network-based identification genetic effect of SARS-CoV-2 infections to Idiopathic pulmonary fibrosis (IPF) patients. Brief Bioinform 2021;22:1254-1266. https://doi.org/10.1093/bib/bbaa235 |
| 200 | Fadista J, Kraven LM, Karjalainen J, Andrews SJ, Geller F, Baillie JK, Wain LV, Jenkins RG, Feenstra B, Initiative C-HG: Shared genetic etiology between idiopathic pulmonary fibrosis and COVID-19 severity. EBioMedicine 2021;65:103277. https://doi.org/10.1016/j.ebiom.2021.103277 |
| 201 | Lee H, Choi H, Yang B, Lee SK, Park TS, Park DW, Moon JY, Kim TH, Sohn JW, Yoon HJ: Interstitial lung disease increases susceptibility to and severity of COVID-19. Eur Respir J 2021;58:2004125. https://doi.org/10.1183/13993003.04125-2020 |
| 202 | Akl MR, Foudah AI, Ebrahim HY, Meyer SA, Sayed KAE: The marine-derived sipholenol A-4-O-3′, 4′-dichlorobenzoate inhibits breast cancer growth and motility in vitro and in vivo through the suppression of Brk and FAK signaling. Mar Drugs 2014;12:2282-2304. https://doi.org/10.3390/md12042282 |
| 203 | Groweiss A, Shmueli U, Kashman Y: Marine toxins of Latrunculia magnifica. J Org Chem 1983;48:3512-3516. https://doi.org/10.1021/jo00168a028 |
| 204 | Kashman Y, Groweiss A, Shmueli U: Latrunculin, a new 2-thiazolidinone macrolide from the marine sponge Latrunculia magnifica. Tetrahedron Lett 1980;21:3629-3632. https://doi.org/10.1016/0040-4039(80)80255-3 |
| 205 | Tsukamoto S, Takeuchi S, Ishibashi M, Kobayashi J: Manzamenones AF from the Okinawan marine sponges Plakortis Sp.: novel dimeric fatty acid derivatives possessing a bicyclo [4.3.0] nonane skeleton. J Org Chem 1992;57:5255-5260. https://doi.org/10.1021/jo00045a048 |
| 206 | Whitson EL, Bugni TS, Chockalingam PS, Concepcion GP, Harper MK, He M, Hooper JN, Mangalindan GC, Ritacco F, Ireland CM: Spheciosterol sulfates, PKCζ inhibitors from a Philippine sponge Spheciospongia sp. J Nat Prod 2008;71:1213-1217. https://doi.org/10.1021/np8001628 |
| 207 | Whitson EL, Bugni TS, Chockalingam PS, Concepcion GP, Feng X, Jin G, Harper MK, Mangalindan GC, McDonald LA, Ireland CM: Fibrosterol sulfates from the Philippine sponge Lissodendoryx (Acanthodoryx) fibrosa: sterol dimers that inhibit PKCζ. J Org Chem 2009;74:5902-5908. https://doi.org/10.1021/jo900844r |
| 208 | Hadjieva P, Popov S, Budevska B, Dyulgerov A: Terpenoids from a Black Sea bryozoan Conopeum seuratum. Z Naturforsch C 1987;42:1019-1022. https://doi.org/10.1515/znc-1987-9-1001 |
| 209 | Sharma S, Guru SK, Manda S, Kumar A, Mintoo MJ, Prasad VD, Sharma PR, Mondhe DM, Bharate SB, Bhushan S: A marine sponge alkaloid derivative 4-chloro fascaplysin inhibits tumor growth and VEGF mediated angiogenesis by disrupting PI3K/Akt/mTOR signaling cascade. Chem Biol Interact 2017;275:47-60. https://doi.org/10.1016/j.cbi.2017.07.017 |
| 210 | Choi YK, Ye BR, Kim J, Kim MS, Lee WW, Ahn GN, Kang N, Jung WK, Heo SJ: Bis (3-bromo-4,5-dihydroxybenzyl) ether, a novel bromophenol from the marine red alga Polysiphonia morrowii that suppresses LPS-induced inflammatory response by inhibiting ROS-mediated ERK signaling pathway in RAW 264.7 macrophages. Biomed Pharmacother 2018;103:1170-1177. https://doi.org/10.1016/j.biopha.2018.04.121 |
| 211 | Choi H, Mascuch SJ, Villa FA, Byrum T, Teasdale ME, Smith JE, Preskitt LB, Rowley DC, Gerwick L, Gerwick WH: Honaucins A−C, potent inhibitors of inflammation and bacterial quorum sensing: synthetic derivatives and structure-activity relationships. Chem Biol 2012;19:589-598. https://doi.org/10.1016/j.chembiol.2012.03.014 |
| 212 | Lin J, Yu J, Zhao J, Zhang K, Zheng J, Wang J, Huang C, Zhang J, Yan X, Gerwick WH: Fucoxanthin, a marine carotenoid, attenuates β-amyloid oligomer-induced neurotoxicity possibly via regulating the PI3K/Akt and the ERK pathways in SH-SY5Y cells. Oxid Med Cell Longev 2017;2017:6792543. https://doi.org/10.1155/2017/6792543 |
| 213 | Zhao D, Kwon SH, Chun YS, Gu MY, Yang HO: Anti-neuroinflammatory effects of fucoxanthin via inhibition of Akt/NF-κB and MAPKs/AP-1 pathways and activation of PKA/CREB pathway in lipopolysaccharide-activated BV-2 microglial cells. Neurochem Res 2017;42:667-677. https://doi.org/10.1007/s11064-016-2123-6 |
| 214 | Satomi Y: Antitumor and cancer-preventative function of fucoxanthin: A marine carotenoid. Anticancer Res 2017;37:1557-1562. https://doi.org/10.21873/anticanres.11484 |
| 215 | Eom SH, Lee EH, Park K, Kwon JY, Kim PH, Jung WK, Kim YM: Eckol from Eisenia bicyclis inhibits inflammation through the Akt/NF‐κB signaling in Propionibacterium acnes‐induced human keratinocyte Hacat cells. J. Food Biochem 2017;41:e12312. https://doi.org/10.1111/jfbc.12312 |
| 216 | Wang S, Wang LJ, Jiang B, Wu N, Li X, Liu S, Luo J, Shi D: Anti-angiogenic properties of BDDPM, a bromophenol from marine red alga Rhodomela confervoides, with multi receptor tyrosine kinase inhibition effects. Int J Mol Sci 2015;16:13548-13560. https://doi.org/10.3390/ijms160613548 |
| 217 | Wang R, Zhang Q, Peng X, Zhou C, Zhong Y, Chen X, Qiu Y, Jin M, Gong M, Kong D: Stellettin B induces G1 arrest, apoptosis and autophagy in human non-small cell lung cancer A549 cells via blocking PI3K/Akt/mTOR pathway. Sci Rep 2016;6:1-10. https://doi.org/10.1038/srep27071 |
| 218 | Lee H, Kim JS, Kim E: Fucoidan from seaweed Fucus vesiculosus inhibits migration and invasion of human lung cancer cell via PI3K-Akt-mTOR pathways. PloS One 2012;7:e50624. https://doi.org/10.1371/journal.pone.0050624 |