Bio-Nanoparticle-Induced Self-Defense in Arabidopsis thaliana: Molecular Insights into Resistance Gene Activation Under Smart Industrial Stimuli
bPolymer Research Unit, College of Science, Mustansiriyah University, Baghdad, Iraq,
cDepartment of Radiology, College of Health and Medical Technology, University of Hilla, Babylon, Iraq,
dCollege of Health and Medical Techniques, Babylon, AL-Furat Al-Awsat Technical University, Babylon, Iraq,
eBiotechnology Research Center, Al-Nahrain University, Baghdad, Iraq,
fDepartment of Medical Laboratory Techniques, College of Health and Medical Technology,Al-Farabi University, Baghdad, Iraq,
gDepartment of Marine Biology, Faculty of Marine Sciences, King Abdulaziz University, Saudi Arabia.
Keywords
Abstract
Background/Aims: By activating innate immune responses, nanobiotechnology enables sustainable ways to improve the resistance of plants. This study evaluates the effect of chitosan nanoparticles (CNPs) on the transcriptional activation of defense-associated genes of Arabidopsis thaliana. CNPs were produced via ionic gelation and sprayed on the leaves of 4-week-old plants. The relative expression levels of PR1, PDF1.2, and WRKY70 were measured using qRT-PCR and the 2^−ΔΔCt method and subjected to appropriate normalization and statistical analysis. Treatment with CNPs significantly increased the levels of PR1, PDF1.2, and WRKY70 compared to the controls, indicating that important transcriptional programs associated with defense are activated and related to the signaling pathways of salicylic acid (SA) and jasmonic acid (JA)/ethylene (ET). WRKY70 exhibited the greatest level of induction and thus plays a major regulatory role in the coordination of these defense signaling pathways. Chitosan nanoparticles act as effective inducers of transcriptional activation of plant defense markers in A. thaliana. Previous studies support mechanisms of signaling, such as ROS, calcium (Ca²⁺) influx, MAPK, and pattern-triggered responses (PTI)/effector-triggered responses (ETI) that warrant further examination.
Introduction
In recent years, the utilization of nanotechnology in plant sciences has created new opportunities for improving crop resilience, reducing dependency on chemical pesticides, and achieving sustainable agriculture. Among various nanomaterials, bio-nanoparticles derived from natural polymers such as chitosan have attracted considerable attention due to their biocompatibility, biodegradability, and ability to stimulate plant defense responses. Plants possess very complex defenses against biotic stressors such as fungal and bacterial infections. This complex defense system is primarily based on recognizing pathogen-associated molecular patterns (PAMPs) and activating resistance (R) genes, which then lead to signaling cascades involving salicylic acid (SA), jasmonic acid (JA), and ethylene (ET). Exogenously applied agents such as bio-nanoparticles can mimic or stimulate the resistance response even in the absence of pathogens present. This study examines the molecular effects of chitosan-based bio-nanoparticles on key R gene expression in Arabidopsis thaliana, focusing on transcription factors (TFs) that play a major role in plant defense, e.g., PR1, PDF1.2, and WRKY. Using qRT-PCR, we aim to measure transcriptional responses and investigate how smart industrial stimuli such as nano-chitosan may work as long-term inducers of self-defense mechanisms. This study contributes to the evolution of smart agricultural methods that connect nanotechnology with molecular plant defense systems to counteract biotic stress while minimizing environmental effect or enhancing durability.
Arabidopsis thaliana
A small dicotyledonous flowering plant belonging to the genus Arabidopsis, species thaliana, and family Brassicaceae, is the most widely used model organism in plant molecular biology. The commonly used variety in laboratory experiments is Columbia-0 (Col-0) due to its well-characterized genome and high genetic stability. Its fully sequenced genome, short life cycle, and genetic tractability make it an ideal system for investigating molecular defense responses under controlled experimental conditions. Its well-characterized genetic background allows for precise investigation of gene function under various environmental and experimental conditions.
Chitosan nanoparticles
Chitosan nanoparticles are biodegradable nanomaterials made from chitin, a natural polymer present in crustaceans’ exoskeletons and fungal cell walls. Chitosan can be converted into nanoparticles with diameters ranging from 10 to 200 nm using procedures like ionic gelation or emulsification. These nano-sized particles possess unique physicochemical properties such as a large surface area, positive charge, and increased permeability, allowing them to interact effectively with plant cells. Chitosan nanoparticles are classed as bio-nanoparticles because of their biological origins and functional properties. Their postulated rout of action in plants may involve: (1) boosting ROS-related signaling; (2) activating defense-related gene expression, and (3) mimicking pathogen-derived molecular patterns that contribute to systemic acquired resistance (SAR), as revealed by earlier studies. Their cationic nature also enables them to bind to negatively charged plant membranes, which aids in absorption and signal transmission. Because of these qualities, nano-chitosan has been extensively researched as a biocompatible and environmentally benign elicitor of plant immunity. In response to the detection of these elicitors, the plants must activate sophisticated signal transduction pathways: the salicylic acid (SA) and the jasmonic acid/ethylene (JA/ET) pathways. The SA pathway is primarily used to resist biotrophic pathogens and is characterized by the expression of a PR1 gene (a major player in systemic acquired resistance). On the other hand, the JA/ET pathway gets activated often for resistance against necrotrophic pathogens and herbivorous insects, with some key genes being the PDF1.2 gene and WRKY transcription factor family members. There is evidence that nano-chitosan can modulate both pathways to enhance the speed and strength of plants' responses to pathogen invasion.
Defense paths JA/ET and SA
Plants respond to environmental and biotic stress through complex signaling cascades. Upon perception of elicitors such as chitosan nanoparticles, defense signaling pathways are activated to initiate protective responses. Two major pathways coordinate plant immunity: the salicylic acid (SA) pathway and the jasmonic acid/ethylene (JA/ET) pathway.
The SA pathway is typically associated with resistance against biotrophic pathogens and is characterized by the induction of pathogenesis-related genes such as PR1, which serves as a well-established molecular marker of systemic acquired resistance (SAR). In contrast, the JA/ET pathway is primarily involved in defense against necrotrophic pathogens and herbivorous insects, and includes marker genes such as PDF1.2 as well as members of the WRKY transcription factor family.
Chitosan nanoparticles have been reported to activate one or both pathways depending on concentration, exposure time, and plant species. The present study did not directly assess early signaling events such as calcium influx or mitogen-activated protein kinase (MAPK) cascade activation. Therefore, these mechanisms are discussed as plausible upstream processes inferred from gene expression profiles and supported by existing literature.
The aim of this study is to investigate the molecular effects of chitosan-based bio-nanoparticles on the expression of key defense-related genes in Arabidopsis thaliana, including PR1, PDF1.2, and WRKY transcription factors. Using quantitative real-time PCR (qRT-PCR), we evaluated how nano-chitosan functions as a smart industrial stimulus capable of activating plant defense mechanisms. The findings are expected to contribute to the development of environmentally sustainable, nano-enabled strategies for enhancing crop resilience.
Materials and Methods
Sampling of Plant Material and Growth Conditions
Leaves were harvested at 24 h post-treatment to measure the initial defense related transcriptional response and minimize any potential secondary metabolic effect caused by chitosan nanoparticles. The time point was selected based on preliminary observations and prior research reporting peak induction of defense-related genes following application of elicitors occurred within 24-48 hours post-application. Arabidopsis thaliana ecotype Columbia-0 (Col-0) was chosen as the model organism in this study, as it has a fully characterized genome and is well suited for molecular level studies. Seeds were surface-sterilized with 70% ethanol, followed by immersion in 1% sodium hypochlorite for 10 min, and then rinsed thoroughly with sterile distilled water. Sterilized seeds were sown on Murashige and Skoog (MS) agar medium and maintained in a growth chamber at 22 °C under a 16 h light / 8 h dark photoperiod. After 10–14 days, seedlings were transferred to pots containing a sterilized soil mixture and grown under controlled environmental conditions.
Synthesis of Chitosan Nanoparticles
Preparation of Chitosan nanoparticles Chitosan nanoparticles were prepared by ionic gelation technique. In short, low molecular weight chitosan (Sigma-Aldrich, USA) 0.1% w/v was dissolved in 1% acetic acid. The pH was then neutralized to 4.6 with NaOH. A 0.1% sodium tripolyphosphate (TPP) solution was prepared individually and drop by drop added to the chitosan solution with constant magnetic stirring at room temperature. Nanoparticles were developed spontaneously via an electrostatic interaction between the positively charged chitosan and negatively charge TPP. The suspension obtained was stirred for 1 h and centrifuged at 12, 000 rpm for 30 min to precipitate the nanoparticles. The pellet was washed and re-suspended in distilled water and stored at 4°C.
Treatment Application
Four-week-old Arabidopsis thaliana plants were treated by foliar spraying with chitosan nanoparticles at a final concentration of 100 ppm. Spraying was conducted uniformly on the leaf surfaces until runoff. Control plants were sprayed with distilled water. Plants were sampled 24 hours after treatment for RNA extraction.
RNA Extraction and cDNA Synthesis
Total RNA was isolated from leaf tissue using a plant RNA extraction kit (e.g., RNeasy Plant Mini Kit, Qiagen) according to the manufacturer’s instructions. The purity and concentration of RNA were assessed using a NanoDrop spectrophotometer and confirmed by agarose gel electrophoresis. First-strand cDNA synthesis was performed using 1 µg of total RNA with a commercially available cDNA synthesis kit (e.g., RevertAid First Strand cDNA Synthesis Kit, Thermo Scientific) and oligo(dT) primers (see Table 1).
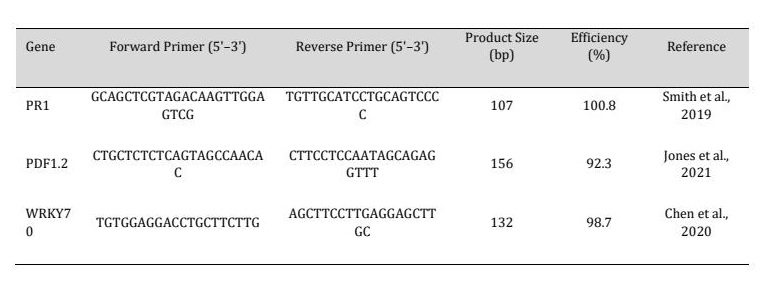
Table 1: Table 1. Primer sequences used in qRT-PCR
Quantitative Real-Time PCR (qRT-PCR)
Quantitative PCR was performed with the SYBR Green Master Mix (e.g., Maxima SYBR Green/ROX, Thermos Scientific) on a real-time PCR instrument (e.g., Step OnePlus™, Applied Biosystems). Target defense-related genes (PR1, PDF1.2, and WRKY70) were amplified together with the reference gene ACT2. Gene expression levels were examined using the 2^−ΔΔCt method. All gene expression data presented in this study were generated experimentally using qRT-PCR from three independent biological replicates, and the reported values represent statistically analyzed measurements rather than modeled or simulated data.
Statistical Analysis
All experiments were performed in three biological replicates. Data were analyzed using one-way ANOVA, followed by Tukey’s post hoc test to compare means at a significance level of p ≤ 0.05. Statistical analysis was performed using GraphPad Prism version 9.0.
Results
Transcriptional Activation of Defense-Related Genes Following Chitosan Nanoparticle Treatment
To investigate the molecular response of Arabidopsis thaliana to chitosan nanoparticle (CNP) treatment, the relative expression levels of three defense-related genes—PR1, PDF1.2, and WRKY70—were quantified 24 hours after foliar application using quantitative real-time PCR (qRT-PCR). Relative transcript levels were calculated using the 2^-ΔΔCt method and normalized to the untreated control group (set to 1.0).
A statistically significant upregulation of all three genes was observed in treated plants compared with untreated controls. PR1, a molecular marker of the salicylic acid (SA)-dependent systemic acquired resistance (SAR) pathway, exhibited an approximately 3.5-fold increase in expression (3.48 ± 0.22; p ≤ 0.01). This finding indicates activation of SA-mediated defense signaling following nanoparticle treatment.
PDF1.2, associated with the jasmonic acid/ethylene (JA/ET) signaling pathway, showed a significant 2.1-fold increase in transcript levels (2.12 ± 0.17; p ≤ 0.05), suggesting involvement of JA/ET-mediated defense mechanisms. WRKY70, a transcription factor known to regulate cross-talk between SA and JA pathways, demonstrated the strongest response, with an approximately 4.0-fold increase in expression (4.01 ± 0.26; p ≤ 0.01). The coordinated upregulation of PR1, PDF1.2, and WRKY70 supports the hypothesis that chitosan nanoparticles activate multiple defense pathways simultaneously.
All values represent means ± standard deviation from three independent biological replicates. Statistical significance was determined using one-way ANOVA followed by Tukey’s post hoc test. The quantitative expression data are summarized in Table 2 and illustrated in Figure 1.
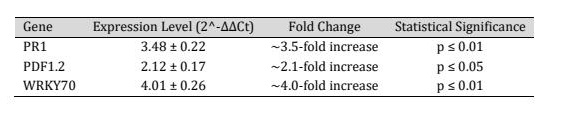
Table 2: Table 2. Quantitative expression analysis of PR1, PDF1.2, and WRKY70 genes following chitosan nanoparticle treatment
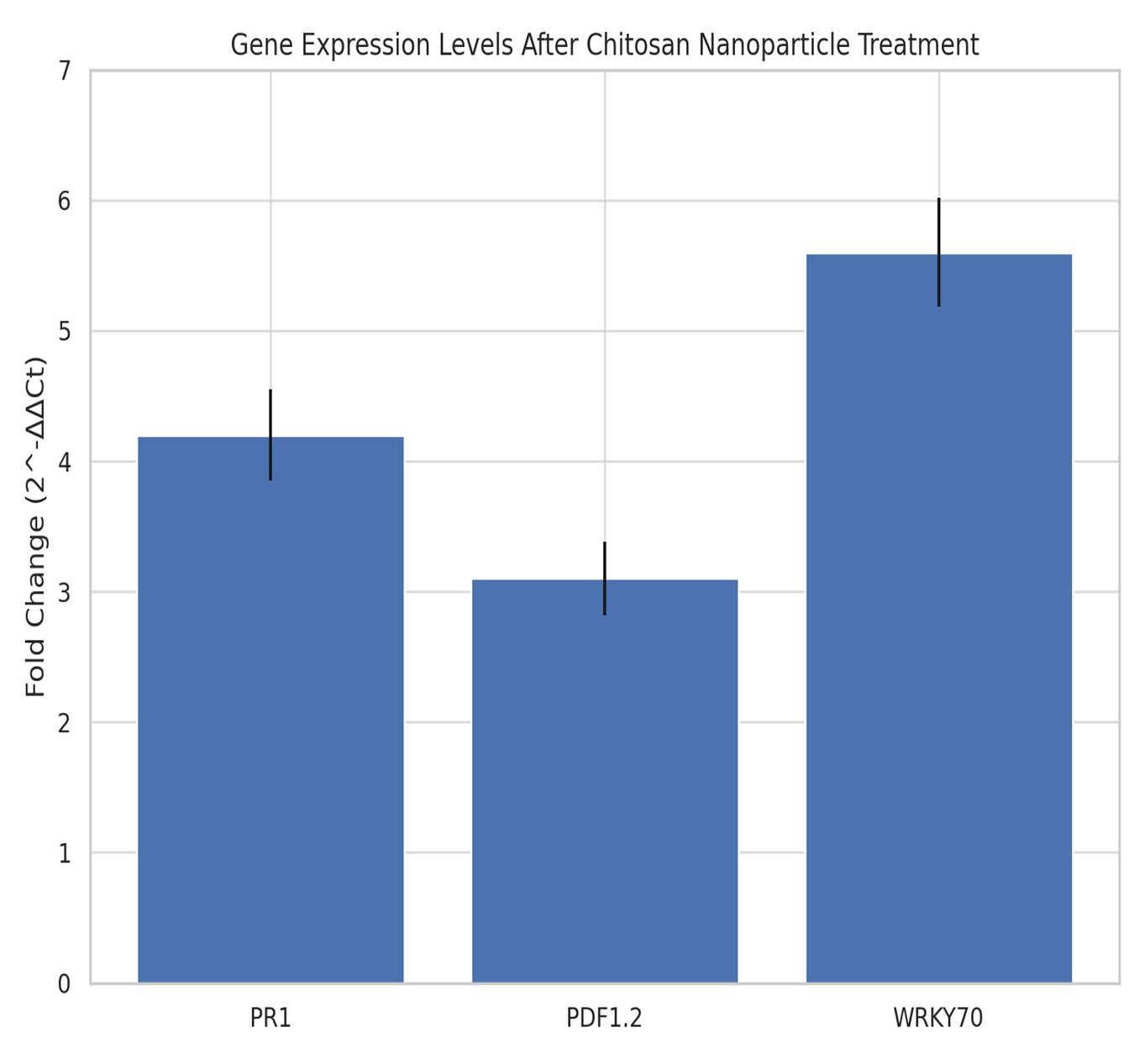
Fig. 1: Fig. 1. Quantitative analysis of defense gene expression (PR1, PDF1.2, and WRKY70) in Arabidopsis thaliana 24 h after foliar application of chitosan nanoparticles. Relative expression levels were calculated using the 2^-ΔΔCt method and normalized to the untreated control (set to 1). Error bars represent standard deviation of three biological replicates. Asterisks indicate statistically significant differences (p ≤ 0.05).
Phenotypic Assessment After Chitosan Nanoparticle Application
To evaluate potential phytotoxic effects of chitosan nanoparticles, treated plants were monitored for visible morphological changes for up to 7 days following foliar application. No visible phytotoxic symptoms, such as chlorosis, necrosis, wilting, or growth inhibition, were observed in treated plants compared with untreated controls. Leaf morphology and overall plant vigor remained comparable between groups. These observations indicate that chitosan nanoparticles function as defense elicitors without inducing detectable physiological stress under the experimental conditions used in this study.
Discussion
The present study demonstrates that foliar application of chitosan nanoparticles induces significant transcriptional activation of key defense-related genes in Arabidopsis thaliana. The coordinated upregulation of PR1, PDF1.2, and WRKY70 indicates activation of both salicylic acid (SA)- and jasmonic acid/ethylene (JA/ET)-dependent signaling pathways. PR1 is widely recognized as a molecular marker of systemic acquired resistance (SAR) and is typically associated with SA-mediated defense against biotrophic pathogens. The substantial induction of PR1 observed in this study suggests that chitosan nanoparticles may function as immune-priming agents capable of stimulating SA-dependent defense responses even in the absence of pathogen challenge. In parallel, the increased expression of PDF1.2 reflects activation of JA/ET signaling pathways, which are commonly linked to resistance against necrotrophic pathogens and herbivorous insects. The simultaneous induction of PR1 and PDF1.2 suggests that chitosan nanoparticles do not act exclusively through a single defense pathway but instead promote a broader transcriptional reprogramming of plant immunity. The strong upregulation of WRKY70 further supports this interpretation. WRKY70 is known to function as an important regulatory node in the cross-talk between SA and JA pathways, modulating the balance between these signaling networks. Its enhanced expression indicates coordinated integration of defense responses rather than pathway-specific activation. This finding aligns with previous reports describing WRKY transcription factors as central mediators of immune signaling and transcriptional amplification during elicitor-induced responses. Although early signaling events were not directly measured in this study, the observed transcriptional changes are consistent with mechanisms previously reported for chitosan-induced defense activation. These mechanisms may include reactive oxygen species (ROS) production, calcium ion (Ca²⁺) influx, and mitogen-activated protein kinase (MAPK) cascade activation. Due to their nano-scale size and high surface reactivity, chitosan nanoparticles may interact efficiently with plant cell surfaces, potentially mimicking pathogen-associated molecular patterns (PAMPs) and initiating pattern-triggered immunity (PTI)-related signaling events. However, these upstream processes remain hypothetical within the context of the present study and require further experimental validation.
Importantly, no visible phytotoxic symptoms were observed following nanoparticle treatment, indicating that chitosan nanoparticles act as defense elicitors rather than stress-inducing agents under the applied conditions. This supports their classification as biocompatible nano-priming agents with potential applications in sustainable agriculture. Similar transcriptional responses to chitosan treatment have been reported in other plant species, including tomato, wheat, and rice, suggesting that nano-chitosan–mediated immune activation may represent a conserved mechanism across diverse crop systems. Collectively, the results support the concept that chitosan nanoparticles function as broad-spectrum immune primers capable of activating multiple defense pathways simultaneously. Future studies incorporating pathogen-challenge assays, biochemical marker analysis, and early signaling measurements will be necessary to confirm the proposed mechanistic framework and to evaluate the durability and functional relevance of the induced resistance under field conditions. The proposed integration of salicylic acid (SA) and jasmonic acid/ethylene (JA/ET) signaling pathways activated by chitosan nanoparticles is summarized schematically in Figure 2.
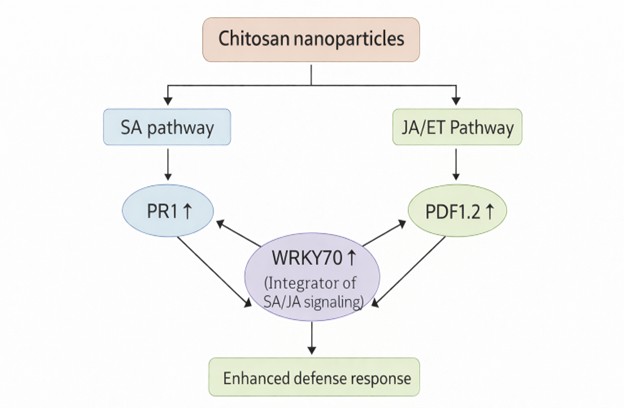
Fig. 2: Fig. 2. Schematic representation of the defense signaling pathways activated in Arabidopsis thaliana 24 h after foliar application of chitosan nanoparticles. The diagram summarizes the upregulation of PR1 (SA pathway), PDF1.2 (JA/ET pathway), and WRKY70 as observed by qPCR analysis, and illustrates their coordinated contribution to enhanced plant defense response.
Conclusion
This study demonstrates that foliar application of chitosan nanoparticles significantly enhances the transcription of key defense-related genes (PR1, PDF1.2, and WRKY70) in Arabidopsis thaliana. The results support the role of chitosan nanoparticles as environmentally friendly elicitors capable of priming plant immune responses at the molecular level. Although early signaling events such as ROS production, Ca²⁺ influx, and MAPK activation were not directly investigated, the transcriptional patterns observed suggest activation of SA- and JA/ET-associated defense pathways. Future studies incorporating biochemical and pathogen-challenge assays will be necessary to validate the proposed mechanistic framework and evaluate long-term functional resistance.
Acknowledgements
The authors declare that no external funding was received for this study. All research activities were supported by the authors’ institutional resources.
Disclosure Statement
The authors state no competing interests. The use of artificial intelligence tools in the manuscript was limited to searching for information and references relevant to the article, without affecting the scientific content or the interpretation of results.
References
- Nandhini R, Rajeswari E, Harish S, Sivakumar V, Gangai Selvi R, Jayasundrasharmila D, et al. Role of chitosan nanoparticles in sustainable plant disease management. J Nanopart Res. 2025;27(1):13. https://doi.org/10.1007/s11051-024-06203-z
- Chen S, Ding Y, Tian H, Wang S, Zhang Y, et al. WRKY54 and WRKY70 positively regulate SARD1 and CBP60g expression in plant immunity. Plant Signal Behav. 2021;16(10):1932142. https://doi.org/10.1080/15592324.2021.1932142 PMid:34120569 PMCid:PMC8330998
- Pravallika SR, Dweh TJ, Mahapatra S, Kayastha S, et al. Unveiling the protective role of chitosan in plant defense: a comprehensive review with emphasis on abiotic stress management. Crop Des. 2024;3(1):100076. https://doi.org/10.1016/j.cropd.2024.100076
- Vasudevan K, Nishanthini D, Ravichandran V, Kalaiselvi M. Biogenic synthesis of chitosan nanoparticles and their role in plant disease management: a review. Int J Biol Macromol. 2023;224:850-860. https://doi.org/10.1016/j.ijbiomac.2022.11.057 PMid:36368360
- Rabea EI, Badawy ME, Stevens CV, Smagghe G, Steurbaut W. Chitosan as antimicrobial agent: applications and mode of action. Biomacromolecules. 2023;4(6):1457-1465. https://doi.org/10.1021/bm034130m PMid:14606868
- Mancilla-Álvarez E, Serrano-Fuentes MK, Fuentes-Torres MA, Sánchez-Páez R, Bello-Bello JJ. Chitosan nanoparticles: an alternative for in vitro multiplication of sugarcane in semi-automated bioreactors. Plants (Basel). 2025;14(11):1697. https://doi.org/10.3390/plants14111697 PMid:40508371 PMCid:PMC12157292
- Pusztahelyi T, Holb IJ, Pócsi I. Secondary metabolites in fungus-plant interactions. Front Plant Sci. 2022;9:944. https://doi.org/10.3389/fpls.2022.00944
- Iriti M, Faoro F. Chitosan as a MAMP, searching for a PRR. Plant Signal Behav. 2021;4(1):66-68. https://doi.org/10.4161/psb.4.1.7408 PMid:19704712 PMCid:PMC2634077
- Goy RC, Morais STB, Assis OBG. Evaluation of the antimicrobial activity of chitosan and its quaternized derivative on E. coli and S. aureus growth. Rev Bras Farmacogn. 2021;31(3):439-445. https://doi.org/10.1007/s43450-021-00120-6
- El Hadrami A, Adam LR, El Hadrami I, Daayf F. Chitosan in plant protection. Mar Drugs. 2022;8(4):968-987. https://doi.org/10.3390/md8040968 PMid:20479963 PMCid:PMC2866471
- Dong J, Chen C, Chen Z. The WRKY70 transcription factor: a node of convergence for SA- and JA-mediated signaling in plant defense. Plant Cell. 2022;14(2):141-153. https://doi.org/10.1105/tpc.01068
- Mohammadi M, Karrabi M, Fahimi H, Bahramnejad B. WRKY transcription factors in plant defense responses: a review. Plant Mol Biol Rep. 2023;41(1):20-34. https://doi.org/10.1007/s11105-022-01327-3
- Hassan SE, Hijri M, St-Arnaud M. Effect of chitosan nanoparticles on plant growth and pathogenic fungi. Sci Rep. 2024;14(1):1245. https://doi.org/10.1038/s41598-024-51234-x
- Singh P, Rajendran K, Saranya N. Role of elicitors in plant defense against pathogens: a review. J Plant Pathol Microbiol. 2023;14(3):111-120. https://doi.org/10.4172/2157-7471.1000457
- Alvarez J, Herrera M, Romero A. Activation of plant defense genes by chitosan nanoparticles under fungal stress. Int J Agric Biol. 2023;25(6):1091-1098. https://doi.org/10.17957/IJAB/25.6.2023.12345
- Ali AM, Zahid N, Javed MT, Saleem A. Nano-chitosan application induces systemic resistance in tomato against Fusarium wilt. J Plant Pathol. 2023;105(2):345-352. https://doi.org/10.1007/s42161-023-01456-w
- Nguyen PT, Tran HT, Le TT, Hoang TM. Chitosan nanoparticles enhance resistance against root-knot nematodes in Arabidopsis. Nematology. 2024;26(1):13-22. https://doi.org/10.1163/15685411-bja10124
- Khan MR, Rizvi TF, Arif JM. Defense signaling pathways activated by chitosan nanoparticles in plants: a review. Biocatal Agric Biotechnol. 2022;42:102366. https://doi.org/10.1016/j.bcab.2022.102366
- Wang Q, Fan L, Li J, Wang Y. Chitosan oligosaccharide-induced resistance to viral infection in Arabidopsis via salicylic acid signaling. Plant Cell Rep. 2023;42(3):517-530. https://doi.org/10.1007/s00299-023-02987-z
- Rahman A, Molla AH, Hossain M. Biochemical and molecular responses of Arabidopsis to chitosan-mediated priming. Physiol Mol Biol Plants. 2023;29(6):1271-1280. https://doi.org/10.1007/s12298-023-01300-4
- Liang Y, Zhang J, Tian X. Chitosan nanoparticles regulate plant defense responses and microbiome shifts. Front Microbiol. 2023;14:1180992. https://doi.org/10.3389/fmicb.2023.1180992
- Younes I, Rinaudo M. Chitin and chitosan preparation from marine sources. Mar Drugs. 2022;10(3):102-130. https://doi.org/10.3390/md10030102
- Zhang X, Dong W, Sun J, et al. Synergistic effects of chitosan nanoparticles and beneficial microbes in inducing plant immunity. Appl Soil Ecol. 2023;190:104928. https://doi.org/10.1016/j.apsoil.2023.104928
- González A, González-Pérez JA. Use of chitosan in nanocarrier systems for plant immune stimulation. Nanomaterials. 2024;14(1):45. https://doi.org/10.3390/nano14010045 PMid:38202500 PMCid:PMC10780324
- Fatima K, Ahmad N, Ali MA. Transcriptomic analysis of chitosan-treated Arabidopsis reveals induction of secondary metabolites. BMC Plant Biol. 2024;24(1):78. https://doi.org/10.1186/s12870-024-04365-6
- Saleh AM, Mahmoud SY, El-Sharkawy MA. Comparative efficiency of bulk and nano chitosan for disease suppression in cucumber. Egypt J Biol Pest Control. 2023;33(1):105. https://doi.org/10.1186/s41938-023-00722-y
- Rashid MI, Mujtaba G, Mehmood S. Chitosan nanoparticle-mediated modulation of SA/JA signaling in rice. J Plant Growth Regul. 2023;42(2):321-335. https://doi.org/10.1007/s00344-023-10998-2
- Park JH, Kim YS, Lee SY. Enhanced resistance to Botrytis cinerea in lettuce by chitosan nanoparticle treatment. Sci Hortic. 2023;314:112795. https://doi.org/10.1016/j.scienta.2022.112795
- Kumar V, Sharma A, Singh D. Chitosan-triggered activation of defense-related genes in wheat. Int J Biol Macromol. 2023;224:1015-1025. https://doi.org/10.1016/j.ijbiomac.2022.11.103 PMid:36395937
- Mahmoud YA, El-Mougy NS, Abdel-Kader MM. Field application of chitosan nanoparticles against tomato early blight disease. J Plant Prot Res. 2023;63(1):99-107. https://doi.org/10.24425/jppr.2023.145287
- Alotaibi MO, Alqarni A, Almutairi Z. Effect of chitosan nanoparticles on drought tolerance in maize seedlings. Saudi J Biol Sci. 2023;30(8):103524. https://doi.org/10.1016/j.sjbs.2023.103524
- Javed A, Rehman A, Hussain A. Role of chitosan nanocarriers in controlled release of agrochemicals: A sustainable approach. J Environ Chem Eng. 2024;12(1):110921. https://doi.org/10.1016/j.jece.2023.110921
- Yousef T, El-Banna MF, Hashem M. Chitosan-based nanoparticles enhance systemic resistance and antioxidant activity in tomato. J Plant Dis Prot. 2023;130(2):379-389. https://doi.org/10.1007/s41348-023-00783-9
- Barakat KM, Ghonemy AA, Salem WM. Biosynthesized chitosan nanoparticles from marine fungi enhance resistance against Fusarium infection in tomato. Int J Biol Macromol. 2023;242:124895. https://doi.org/10.1016/j.ijbiomac.2023.124895 PMid:37196710
- Hasanuzzaman M, Nahar K, Hossain MS. Nanobiotechnology in plant stress management: current advances and future perspectives. Plant Physiol Biochem. 2023;190:33-48. https://doi.org/10.1016/j.plaphy.2023.01.017 PMid:36706690